Soichi Hayashi
Soichi Hayashi
One of our user ran into a similar issue with TractSeg 2.2. ``` + Tractometry -i tractseg_output/TOM_trackings/ -o tractseg_output/Tractometry_peaks.csv -e tractseg_output/endings_segmentations/ -s tractseg_output/peaks.nii.gz --tracking_format tck --TOM tractseg_output/TOM --peak_length Matplotlib created...
It finished successfully with the current master branch!  Do you think I should try 2.3 branch also? Or is this fixed on latest master branch code?
I am also seeing a very slow streamline loading performance for tck as well. It looks like this part of the code is severely CPU bounded > https://github.com/nipy/nibabel/blob/master/nibabel/streamlines/array_sequence.py#L32
I did a bit more troubleshooting, and I found that if I load plotly like following. ``` import Plotly from 'plotly.js/dist/plotly.min.js' ``` .. the 3d graphs works. But not this...
I *think* I had to add ify-loader (whatever that is..) to my webpack config. build/webpack.base.conf.js under module.exports.module.rules ``` //plotly needs ify-loader (rest should be babel-loader - for es6 syntax) {...
@MiltonCobo you should use https://github.com/David-Desmaisons/vue-plotly instead. It just works.
I believe the validator works.. as I see the error message displayed here > https://openneuro.org/datasets/ds002735/versions/1.0.1/file-display/acq-moldOFF_T1w.json 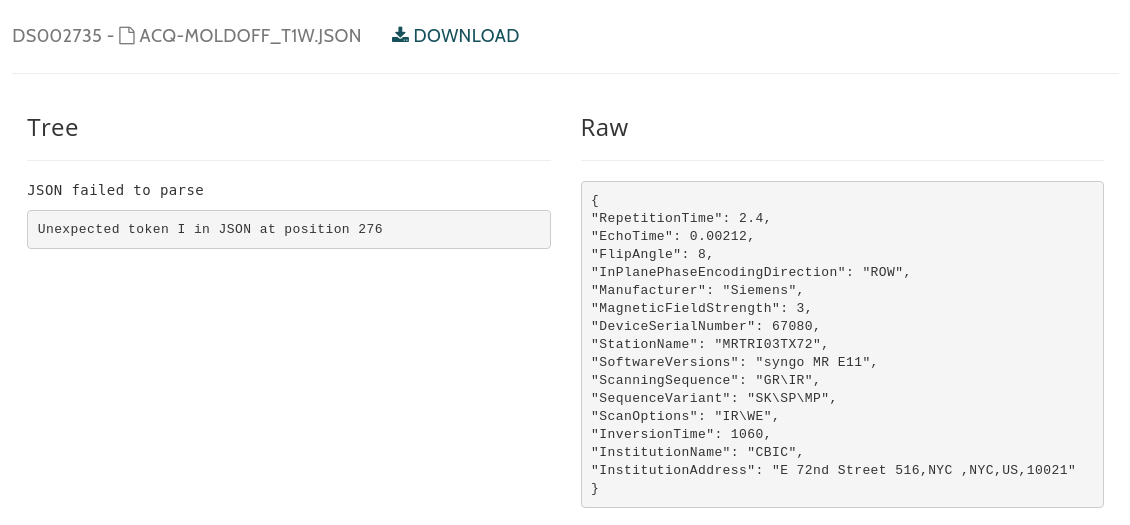 Should OpenNeuro be publishing these files given they are *known* to be invalid? I...
FYI. I am still seeing some parse errors on some of the openneuro published datasets ``` (node:1091) DeprecationWarning: collection.ensureIndex is deprecated. Use createIndexes instead. 2020-08-04T04:07:37.297Z [undefined] [31merror [39m: read ECONNRESET...
How should I handle invalid JSON files that are already published in the OpenNeuro? Is there a plan to purge them from the OpenNeuro, or should I come up with...
@franklin-feingold Yes, it looks like those are the only problematic files that we are finding.