Spatial graph function
Here's that spatial_graph function I had mentioned @giovp and @Mirkazemi.
I figure it should probably be named something different to make sure it's clear it would only work for hex wells.
It looks great!
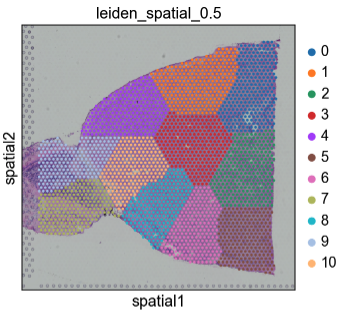
One first improvement could be to expose a parameters that explicitly ask for the number of rings in the neighbors (first 6 ring etc.). Btw, how do I push to this specific PR from my local repo? I managed to pull it but can't figure out a way to push here and not on my fork.
Oh, sorry, I had completely missed your comment here!
It looks great!
Thanks! Can I ask why you used leiden clustering on this?
One first improvement could be to expose a parameters that explicitly ask for the number of rings in the neighbors
This should be easy enough. I'm curious as to whether this it's better to leave this up to whatever algorithm is being used however, since the one step graph has some nice properties. It'd probably be important to include distance in the multistep graph.
Btw, how do I push to this specific PR from my local repo?
This should be fairly straight forward. If you're using the github cli, I think it should just be:
gh pr checkout 1383
# whatever changes
git push
Let me know if that gives you errors, since it might be a repository permissions issue.
@ivirshup @giovp i have updated the function, now it can process non-visium coordinates and have the number of rings option for visium. https://github.com/theislab/spatial-tools/blob/graph/notebooks/build_spatial_graph.ipynb
That's great, I'll add Isaac to that project so he can see code (repo is private). Let's discuss whether to integrate in scanpy at next meeting! Thank you Sergei !
Hey! I was just getting back to this, and I'm not sure I agree with all the choices made in the updates. For example, I would probably rather have separate functions for different spatial neighbor strategies. Also this won't work if coord_type isn't "visium".
Should I be making changes to this PR, or are you relying on it?
hexagonal connectivity
I would propose a separate function just for visium data, maybe called visium_connectivity. It works with the assumption of a hexagonal grid. This removes the neigh, radius, and coord_type arguments.
I think if we have an argument for n_rings there should be some way to weight the connectivity graph by how many steps away each extra point is. Also I'm pretty sure the current implementation of n_rings is incorrect when n_rings>2.
Without weighting, I think it should be more like this:
def walk_nsteps(adj, n):
"""Expand adjacency matrix adj by walking out n steps from each node."""
adj = adj.astype(bool)
cur_step = adj
result = adj.copy()
for i in range(n):
cur_step = adj @ cur_step
cur_step.setdiag(False)
result = result + cur_step
return result
An example showing this works
import networkx as nx
from scipy import sparse
from matplotlib import pyplot as plt
import numpy as np
def walk_nsteps(adj, n):
"""Expand adjacency matrix adj by walking out n steps from each node."""
adj = adj.astype(bool)
cur_step = adj
result = adj.copy()
for i in range(n):
cur_step = adj @ cur_step
cur_step.setdiag(False)
result = result + cur_step
return result
# Test data (path graph)
G = nx.Graph()
G.add_nodes_from([0,1,2,3])
G.add_edges_from([(0, 1), (1, 2), (2, 3), (3, 4)])
adj = nx.adjacency_matrix(G).astype(bool)
fig, axes = plt.subplots(nrows=3)
# Fixed circle layout
pos = {i: (np.cos(-np.pi + (np.pi * i) / 4), np.sin(-np.pi + (np.pi * i) / 4)) for i in range(5)}
for n, ax in enumerate(axes):
nx.draw(
nx.Graph(walk_nsteps(adj, n)),
pos=pos,
ax=ax,
)
plt.show()
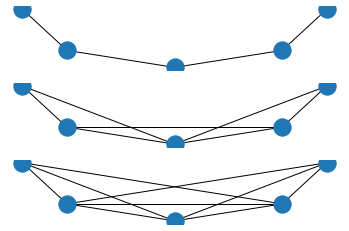
Radius/ n-neighbors coordinates
I'm a little less sure about this use case. Since cells come in all shapes and sizes, I'm not sure if representing each cell as a centroid with a fixed radius or number of neighbors is the way to go. That said, maybe it is.
This could be a separate function spatial_connectivity? In this case, I think it would also be useful to return a distance matrix.
About adding all powers of adjacency matrix - i implemented it at first as you did, but then i thought that it was redundant and changed to the present variant. My thought was that the hexagonal connectivity structure would allow to get all paths of less than n_rings with only n_rings power. And this works in practice, but i agree that there can be some edge cases with isolated node blocks.
@Koncopd here's an example of the current implementation breaking down at n=3
def walk_nsteps_current(adj, n=1):
adj = adj.copy()
if n > 1:
# get up to n_rings order connections
adj += adj ** n
adj.setdiag(0)
adj.eliminate_zeros()
adj.data[:] = 1.0
return adj
fig, axes = plt.subplots(nrows=3)
# Fixed circle layout
pos = {i: (np.cos(-np.pi + (np.pi * i) / 4), np.sin(-np.pi + (np.pi * i) / 4)) for i in range(5)}
for n, ax in enumerate(axes):
nx.draw(
nx.Graph(walk_nsteps_current(adj, n + 1)), # making sure we start at 1
pos=pos,
ax=ax,
)
plt.show()
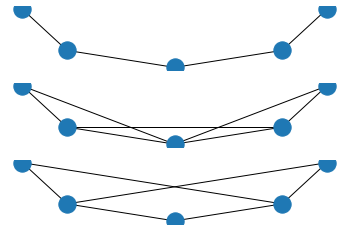
For n=3 you're missing the neighbors at depth=2. Basically, you're just jumping to the 3rd step, instead of accumulating neighbors up to that step.
Yes, but if we have hexagonal connections, then, for example, n-1, n-2 and so on paths will be counted.
2 examples
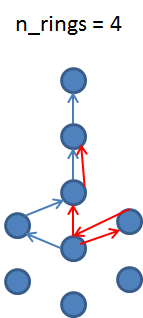 You could check also the notebook i posted above.
You could check also the notebook i posted above.
Just reading along.... if all you want is to find neighbors within a certain number of hops, then non-zero values of powers of the adjacency matrix is a bit inefficient i think. There should be simple breadth-first-search or depth-first-search algorithms implemented in networkx I imagine. And if you're bent on this approach, adding self-loops (diag = 1) will mean you can just do powers.
I'm sorry but I'm not sure I'm following you guys in the discussion, although from the figure it does seem that @Koncopd implementation is missing the n=2.
Regarding radius, this is definitely super useful for anything that is not a grid, and also agree that a distance matrix can be useful, maybe then creating a graph by binarizing that distance matrix based on a threshold?
@LuckyMD thanks for your comment. So it seems that it is enough to do this
adj.setdiag(1) # or 2?
adj += adj ** n
adj.setdiag(0)
adj.eliminate_zeros()
adj.data[:] = 1.0
And this should eliminate all edge cases.
Not sure if it is a good idea to use networkx for this.
@Koncopd yes, I believe that should cover everything (maybe test to make sure I'm not missing sth here). However, I still think taking adjacency matrix powers will not be as fast as a BFS/DFS.
@LuckyMD matrix multiplication on sparse matrices is actually a pretty efficient version of breadth first search, as used by graphBLAS. Here's an example for a BFS from a specific point (which you can expand to all points by using the identity matrix):
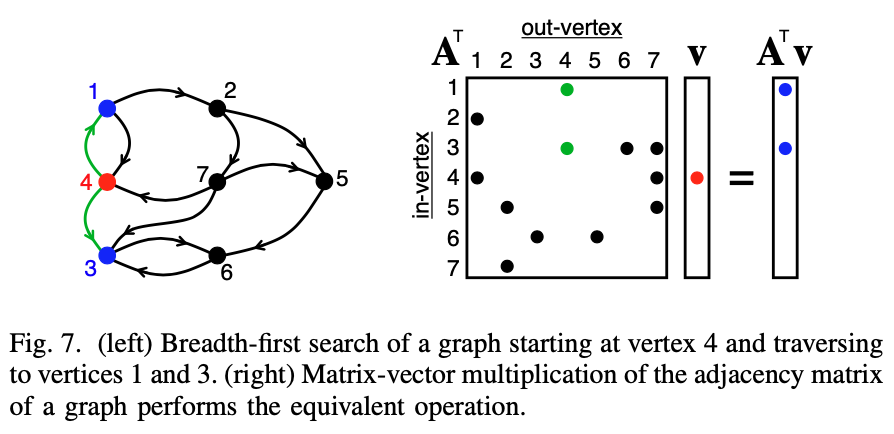
@Koncopd, I think that should work, since you're adding a self edge so you have redundancy at each step.
A separate point of contention on handling it like this: do you want the first step neighbors to have the same weight as the second step neighbors? My assumption would be no.
Btw, I've separated the visium and non-visium case (since they don't share much code), added tests for the visium case, and removed the warnings. I've rebased so I could re-use some test data from another PR.
@LuckyMD matrix multiplication on sparse matrices is actually a pretty efficient version of breadth first search, as used by graphBLAS.
Hmm... i had no idea it was more efficient than using queues, but Figure 10 suggests otherwise. Matrix multiplication definitely seems easier.
do you want the first step neighbors to have the same weight as the second step neighbors?
You could keep these separate by binarizing the adjacency matrix before doing the multiplication. The neighbors that are only reachable via the Nth-hop should always have a 1 in the N-th matrix product that way.
You could keep these separate by binarizing the adjacency matrix before doing the multiplication. The neighbors that are only reachable via the Nth-hop should always have a 1 in the N-th matrix product that way.
Thats a problem with this strategy. Since it's an undirected graph, a node is it's own second neighbor. Since it's a hexagonal grid my first neighbors are also my second neighbors as well as my nth neighbors (in most cases, if there are edges or missing cells this might not be the case). We would either have to do our own BFS which precludes back tracking (i.e. for each search from each node remove previously visited edges), or we could take the difference of the edge sets at each update. Taking the difference would probably be easier.
With self-loops a node is also its first neighbor. That means if you binarize the adjacency matrix and you have a 1 in the adjacency matrix after taking some power, that means there is only 1 way to get to that neighbor and it must thus only be reachable in the N-th hope (N being the power you just multiplied with). You could therefore also just look for the 1s in the matrix after every multiplication.... that might be a bit easier than taking the difference.
Just checked this, and some of the members of the nth ring have more than one neighbor in the nth-1, so checking for values of one doesn't quite work. Also I've been collecting some functions for set operations on sparse indices so taking the difference was quite easy 😉
Here's the check:
Code
import scanpy as sc
from sparse_wrapper._core.coordinate_ops import difference_indices
import numpy as np
from scipy import sparse
def find_steps_val(adj, n_steps):
"""Finding positions with value of 1."""
cur = adj.astype(int)
diffs = [adj.copy()]
for i in range(n_steps):
cur.data[:] = 1
cur = adj @ cur
diffs.append(cur == 1)
return diffs
def find_steps(adj, n_steps):
"""Finding differences in sparsity patterns."""
diffs = [adj.copy()]
prev = adj + sparse.eye(adj.shape[0])
for i in range(n_steps):
cur = prev @ adj
diffs.append(difference_indices(cur, prev))
prev = cur
return diffs
adata = sc.datasets.visium_sge("V1_Adult_Mouse_Brain")
sc.pp.visium_connectivity(adata)
adj = adata.obsp["spatial_connectivity"]
N = 3
diff_res = find_steps(adj, N-1)
val_res = find_steps_val(adj, N-1)
for i in range(N):
adata.obs[f"val_res_{i}"] = np.ravel(val_res[i][50, :].toarray()).astype(int)
adata.obs[f"diff_res_{i}"] = np.ravel(diff_res[i][50, :].toarray()).astype(int)
sc.pl.spatial(
adata,
color = [f"val_res_{i}" for i in range(N)] + [f"diff_res_{i}" for i in range(N)],
ncols=N
)
This is a plot of the neighbors of the 50th well at steps 1, 2, and 3 from each method. Top is finding positions that equaled 1, bottom is taking the differences of the sparsity patterns.
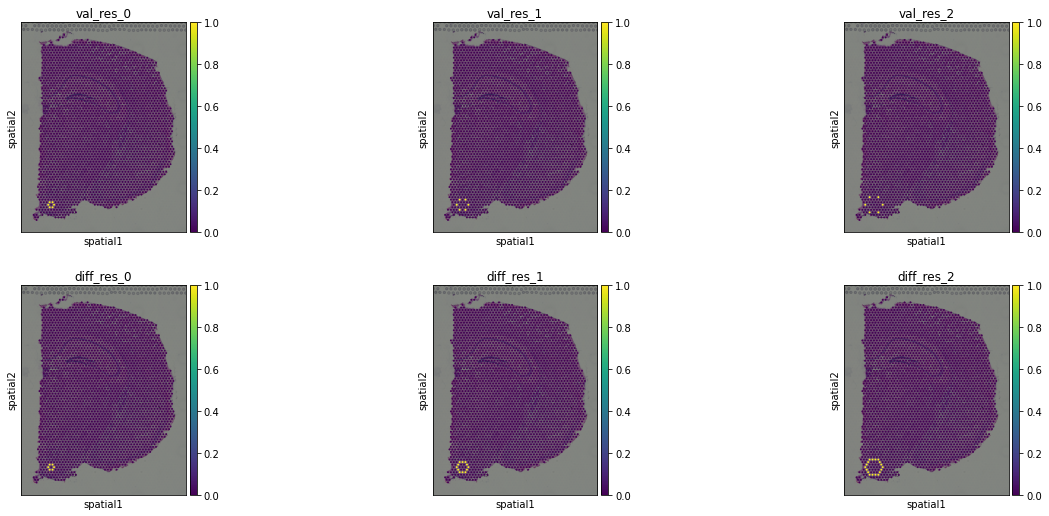

Either way, these look kinda pretty:
from operator import add
from functools import reduce
cells = np.random.choice(adata.n_obs, 20, replace=False)
adata.obs["fireworks"] = (
reduce(add, (diff_res[i][cells] * (3 - i) for i in range(3)))
.max(axis=0)
.toarray()
.ravel()
)
sc.pl.spatial(adata, color="fireworks", cmap="inferno")
some of the members of the nth ring have more than one neighbor in the nth-1
Damn... yeah, that won't work then. Back to differences of adjacency matrices then i guess.
It looks as though your functions are replicating what I assume is going on in the backend of networkx anyway. Are you trying to avoid the heavy dependency or why replicate the effort? I would assume that once we need things like label propagation (maybe to denoise label assignment after deconvolving spatial spots?) then networkx might come in handy, no?
It looks as though your functions are replicating what I assume is going on in the backend of networkx anyway.
So that repo is actually me trying to make it so we can have CSR and CSC matrices that act the same as numpy ndarrays. Those functions are just some building blocks for broadcast operations on those arrays, which are otherwise implemented in C++ in scipy.
A lot of this could probably be done with networkx. networkx is just very slow, since it's all pure python.
A lot of this could probably be done with networkx. networkx is just very slow, since it's all pure python.
Ah, I wasn't aware of that. Cool... then just trying to work with arrays I guess. I hope using spatial graphs doesn't require more than some basic operations in that case.
quick practical comment on this very interesting discussion. @ivirshup shall we have this here or moved to the other package under development? We need to take a decision on this because we'll have to see how it works with rest of functions. Pinging @Koncopd as well. Sorry to put pressure but we are on a tight schedule 😅
re: networkx discussion, also agree with Isaac on having these operations external to networkx.
I'd like to add this function plus tests to scanpy. I think I'll leave out n_rings argument and the radius_neighbors functions until there are clear use-cases. I would recommend just having a copy of the code in spatial-tools, which can be deduplicated later.
ok that's great, thank you! The radius_neighbors have very clear applications in fish-like data, and we are assmebling a tutorial to show exactly that. The n-rings as well especially in the context of cell-cell communication (although did not check that systematically yet). So shall we add this functionality to spatial-tools ?
I could probably be convinced to include radius_neighbors. I'd mainly need a test case. My initial opposition was that (1) it's a pretty trivial implementation and (2) without a real example I'm not sure if it's missing any obvious edge cases.
For n-rings, I think that walking some steps from each node generalizes beyond graph construction, and might be reasonable to have as a separate method. I also haven't seen any recommendations about how to weigh the edges, which I think is pretty important.
Yeah the edge weighing is definitely not clear we would've to test that. ok so only tests are missing ?
ok, after talking with @davidsebfischer we definitely want to have both radius as well as n_rings and also have graph transform functions so to weight edges differently (based on different kernels). I'd suggest to keep the function in spatial-tools and then decide what to port to scanpy.
@ivirshup regarding use cases (for radius, n_rings, and edge weighing) there are multiple ones from graph convolutional networks application, but it could be interesting also just for the other graph analysis that we have implemented in spatial-tools. We'd have to do extensive interpretation of results of course to prove that it really is useful.
super interesting discussion btw. I think this can be closed