feat(core): implement nep13 and nep18 to allow sklearn integration
This is a WIP on having the vaex dataframe and expressions behave more like 2d and 1d numpy arrays. This allows us to feed sklearn's fit a vaex dataframe, and have transform also return a dataframe with virtual columns. This gives us
- A better API for vaex dataframes, since it's more like numpy
- Leverage the sklearn algorithms, and turn them into vaex expressions/virtual columns
- JIT the result of sklearn transforms, so we can use CUDA or numba to optimize them.
- Out of core support for sklearn, since we don't keep anything in memory, it's all streaming algos (at least for the transformers tested here).
Numpy NEPs:
- https://numpy.org/neps/nep-0018-array-function-protocol.html
- https://numpy.org/neps/nep-0013-ufunc-overrides.html
Example usage (5x faster):
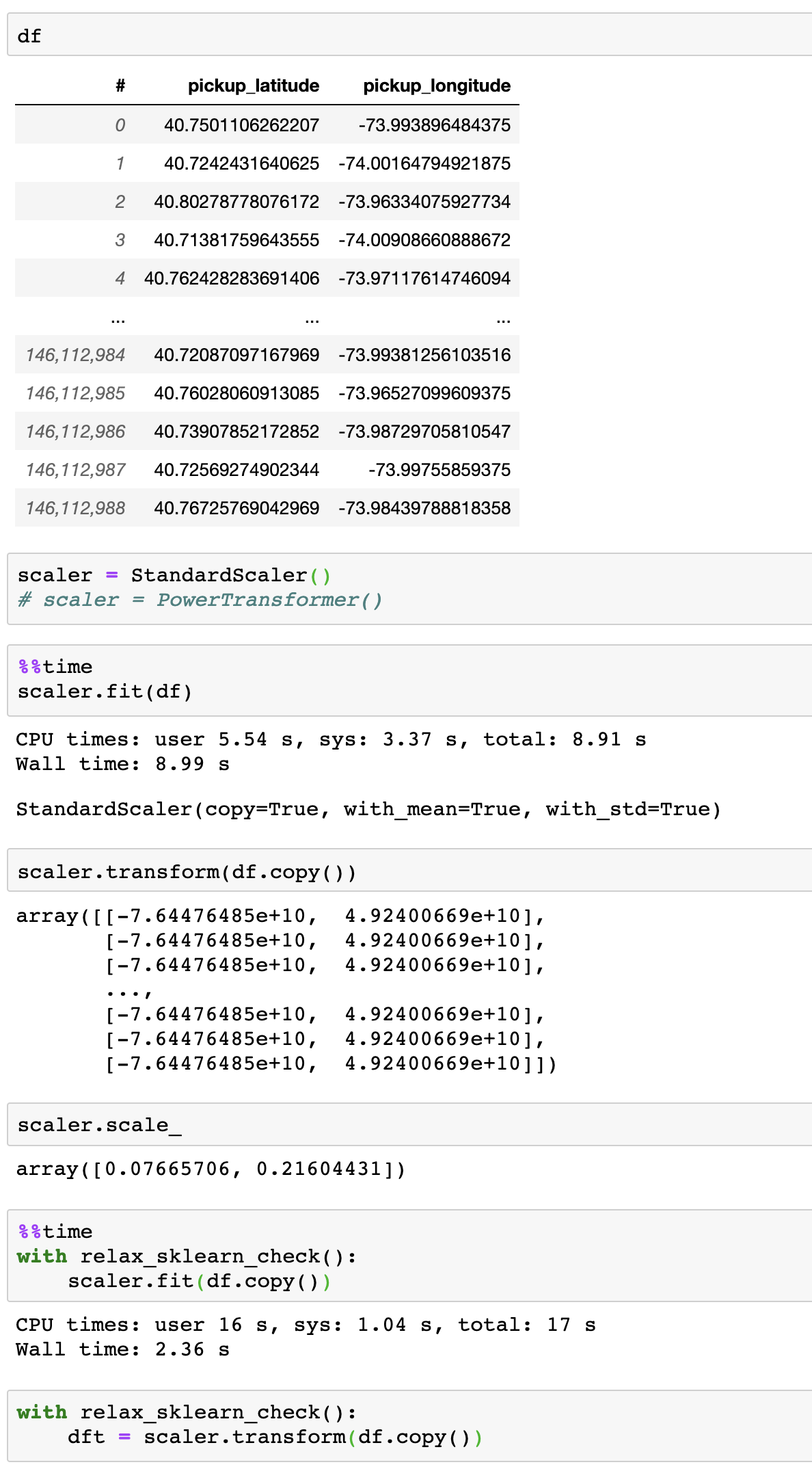
TODO
- [ ] increase coverage of numpy function (for instance to allow PCA?)
- [ ] Test coverage
- [ ] See if we can get sklearn's check_array to allow a vaex dataframe to pass through.
Something that is probably missing is right now is how strings are handled in this scenario, especially with interaction to scikit-learn. For instance:
import vaex
from sklearn.preprocessing import LabelEncoder, OrdinalEncoder
df = vaex.from_arrays(x=['dog', 'dog', 'cat'],
y=['white', 'golden', 'black'])
with sklearn_patch(), df.array_casting_disabled():
enc = OrdinalEncoder()
enc.fit_transform(df[['x', 'y']])
The same happens when using LabelEncoder. Can we support this at all?
I was gonna write this as a unit-test but I was not sure how to fold it in with the rest of the tests..
As this PR is heavily geared towards scikit-learn compatibility, currently the ColumnTransformer does not seem to work (but the Pipline does!). Here is a simple example:
import vaex
import numpy as np
from sklearn.decomposition import PCA
from sklearn.preprocessing import PolynomialFeatures
x = np.random.random_sample(size=10)
y = np.random.randint(0, 15, size=10)
df = vaex.from_arrays(x=x, y=y)
pca = PCA(n_components=2, svd_solver='full', random_state=42)
poly = PolynomialFeatures(degree=2)
# transformers -> List of (name, transformer, column(s)
# Option 1: specify columns by names
col_trans = ColumnTransformer(transformers=[
('pca', pca, ['x', 'y']),
('polynomials', poly, ['x', 'y']),
])
with sklearn_patch(), df.array_casting_disabled():
trans = col_trans.fit_transform(df)
So this will not work because ColumnTransformer expects a pandas DataFrame if the columns are specified with names.
Alternatively one can use column indices:
col_trans = ColumnTransformer(transformers=[
('pca', pca, [0, 1]),
('polynomials', poly, [0, 1]),
])
with sklearn_patch(), df.array_casting_disabled():
trans = col_trans.fit_transform(df)
And this will fail for a different reason.
Or perhaps you have imagined the usage to be bit different and we should not rely on ColumnTransformer. This is absolutely fine with me! Just that the intended usage should be described in some detail somewhere.
For reference, creating a pipeline like this works just fine! :
pca = PCA(n_components=2, svd_solver='full', random_state=42)
poly = PolynomialFeatures(degree=2)
pca_ply_pipe = Pipeline(steps=[('pca', pca),
('polynomial_features', poly
])
with sklearn_patch(), df.array_casting_disabled():
trans = pca_ply_pipe.fit_transform(df)
But overall i really really like the direction this is going :)