find.cluster not clustering groups
Hello,
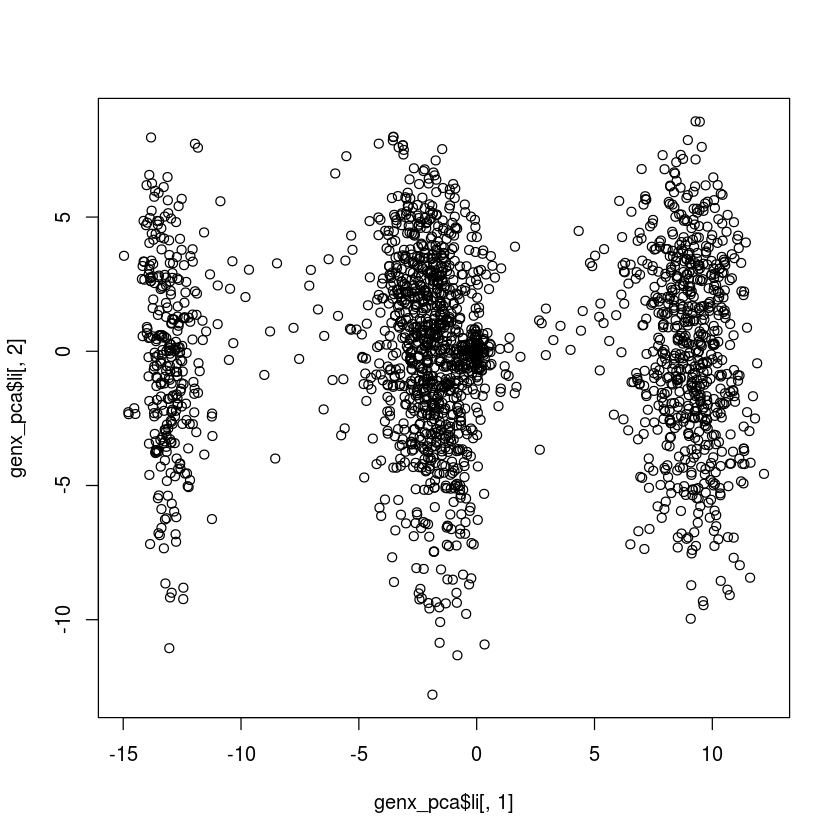
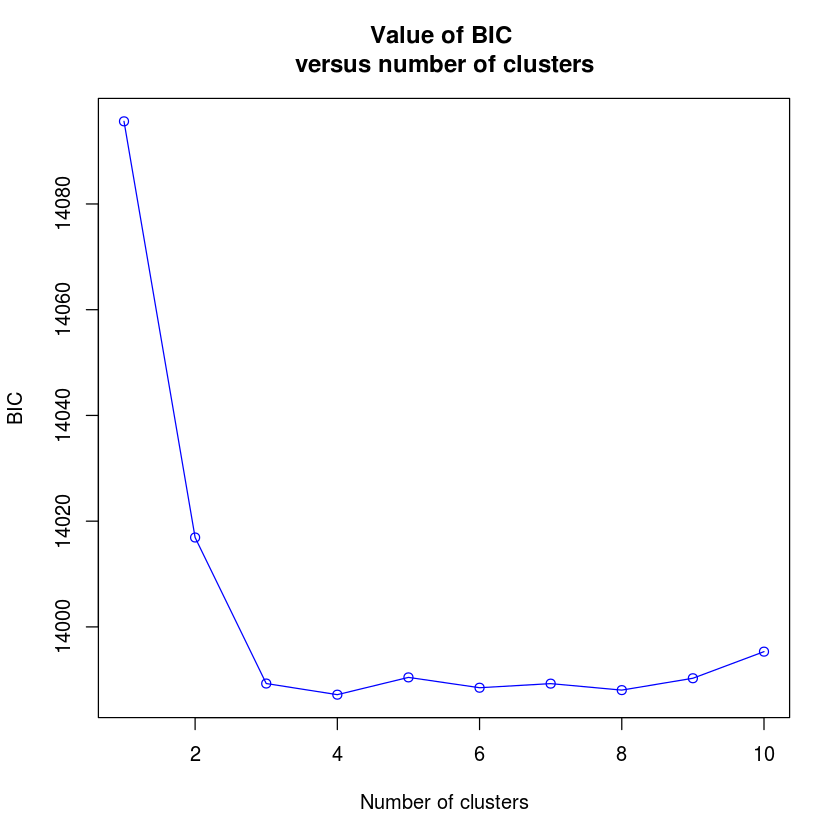
I'm new to adegenet, but I'm following the DAPC tutorial after finding mysterious clustering in a dudi.pca() of my genind format SNP data (747 loci, 2061 individuals). My data appears to have three distinct clusters, and when I run find.clusters, I see k of 3-8 is most appropriate. However, grp$grp yields 1 group with all of my 2061 individuals. I have no informative pop data for this data set, so I've set dat@pop=NULL. The information in grp seems to be wrong, and when I run dapc() I get the error:
Error in svd(X, nu = 0L): infinite or missing values in 'x' Traceback:
- dapc(dat_genx, group$grp)
- dapc.matrix(dat_genx, group$grp)
- dapc(as.data.frame(x), ...)
- dapc.data.frame(as.data.frame(x), ...)
- lda(XU, grp, tol = 1e-30)
- lda.data.frame(XU, grp, tol = 1e-30)
- lda(structure(data.matrix(x), class = "matrix"), ...)
- lda.matrix(structure(data.matrix(x), class = "matrix"), ...)
- lda.default(x, grouping, ...)
- svd(X, nu = 0L)
- stop("infinite or missing values in 'x'")
There is no missing data in my genind object, so it's unclear to me where this is coming from. I have also tried to set arbitrary dat@pop values based on the site where I collected the samples, but even with a populated dat@pop field the find.clusters()/dapc() outputs still have the same issues. Any help would be appreciated, I've combed the adegenet documentation and forum thoroughly.
Thank you very much! Katrina