`find_and_sort_coord` function handles empty-array input errors in an unclear way
Error reported by user on the forum: https://forum.spinalcordmri.org/t/valueerror-when-using-register-to-template/867/2

Will update with more information if/when the user provides an image to try to reproduce the issue with.
I suspect the segmentation is completely empty. If the user is not comfortable sharing the image, we could ask them to share the segmentation so we could at least verify this hypothesis.
From the error message, we know that the issue is due to z_mean being an empty array.
z_mean is created here:
https://github.com/spinalcordtoolbox/spinalcordtoolbox/blob/0bfc2da2eb4c2b93fa70b3b2f24d589b1932489c/spinalcordtoolbox/centerline/core.py#L106-L107
NB: It's worth nothing that here,
im_segis just the image that was passed tosct_register_to_template -s, without any sort of resampling/preprocessing.
Taking a look at the find_and_sort_coord() function, I found its code to be quite difficult to understand.
So, as an exercise, I tried rewriting the function such that each step matches the comments/pseudocode a little better:
# Get [X,Y,Z] coordinates of all non-null values in the segmentation image (==> 3xN array)
data = img_seg.data
coord_nonnull = np.array(data.nonzero())
# Get the dimension (0,1,2) corresponding to the SI axis
dim_si = [img_seg.orientation.find(x) for x in ['I', 'S'] if img_seg.orientation.find(x) != -1][0]
# Get a sorted set of non-empty SI slice numbers (==> list of size M, e.g. [23,24,25...])
si_slices_nonnull = sorted(set(coord_nonnull[dim_si]))
# Get the non-null coordinates within each non-empty slice (==> list of M arrays of size 3xn)
coord_nonnull_per_si_slice = [coord_nonnull[:, coord_nonnull[dim_si] == s] for s in si_slices_nonnull]
# Average the non-null coordinates within each non-empty slice (==> Mx3 array)
coord_average_per_si_slice = [coord_list.mean(axis=-1) for coord_list in coord_nonnull_per_si_slice]
return np.array(coord_average_per_si_slice).T # .T --> (Mx3 array ==> 3xM array)
Sanity check against sct_testing_data
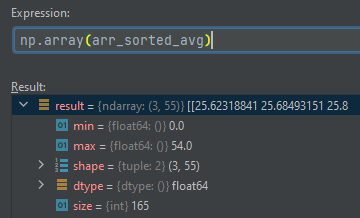
Output of the previous version:
Output of my version:
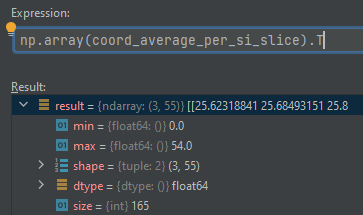
Note: I noticed on the
np.where()docs, it says:
numpy.where(condition, [x, y, ]/)When only
conditionis provided, this function is a shorthand fornp.asarray(condition).nonzero(). Using nonzero directly should be preferred, as it behaves correctly for subclasses.So, that's why I replaced
np.array(np.where(img.data))withdata.nonzero().
The rewritten function suggests that if z_mean is an empty array, then the input segmentation must also be empty, I think? (i.e. there are no non-empty slices in the segmentation)
The main problem on our end, then, is that the find_and_sort_coord function didn't catch the error in a graceful way.
I suspect the segmentation is completely empty. If the user is not comfortable sharing the image, we could ask them to share the segmentation so we could at least verify this hypothesis.
Oh! I missed your reply... I should have refreshed the page, hehe. You came to the same conclusion, but much earlier than I did. :P
Given that I'm fairly confident now that this is non-urgent, I'm going to remove this from the 5.6 milestone, and proceed with the 5.6 release (i.e. generating the 5.6 Changelog)
thank you for the thorough investigation! this is elegant detective work
A quick and easy way to solve this would be to catch an empty input seg and throw an error if that happens