Label transfer increase resolution
Hi, thank you so much for making Seurat available, I am trying to transfer the labels from a 10x dataset to a sciRNA-seq3 dataset.
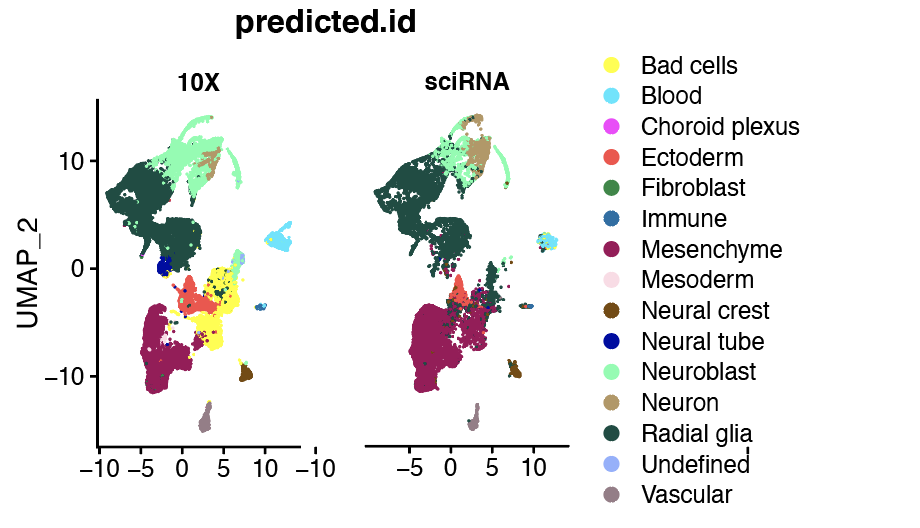
Overall it performs ok, but as you can see in the following figure, the "Bad cell" type disappeared in the sciRNA-seq3. Would you have any idea why that would be ? This Bad cells cluster is lower complexity libraries (low UMI ocunt) that usually get filtered out, but I want to check this cluster across datasets.
Because it takes quite a lot of computational time to re-run everything and optimise the different parameters in the functions, I wanted to know if you had some intuition as to what I could change to have a better resolution ? Thanks a lot for your help , I paste the code hereunder: obj.list <- SplitObject(obj, split.by = "data") for (i in names(obj.list)) { obj.list[[i]] <- SCTransform(obj.list[[i]], verbose = FALSE) }
obj.features <- SelectIntegrationFeatures(object.list = obj.list, nfeatures = 3000) obj.list <- PrepSCTIntegration(object.list = obj.list, anchor.features = obj.features)
reference_dataset <- which(names(obj.list) == "10X")
obj.anchors <- FindIntegrationAnchors(object.list = obj.list, normalization.method = "SCT", anchor.features = obj.features, reference = reference_dataset) obj.integrated <- IntegrateData(anchorset = obj.anchors, normalization.method = "SCT")
obj.integrated <- RunPCA(object = obj.integrated, verbose = FALSE) obj.integrated <- RunUMAP(object = obj.integrated, dims = 1:30)
Idents(obj.integrated) <- [email protected]$data
anchors <- FindTransferAnchors(reference = obj.subset, query = obj.integrated, reduction="rpca", k.anchor=10, reference.reduction)
predictions <- TransferData(anchorset = anchors, refdata = obj.subset$Class)
obj.integrated <- AddMetaData(object = obj.integrated, metadata = predictions) DimPlot(obj.integrated, group.by = "predicted.id", split.by="data", cols=colours)
Hi,
It seems that you need to set normalization.method = "SCT" in FindTransferAnchors .
Do you use SCTransform to normalize your reference, obj.subset?