qca-dataset-submission
 qca-dataset-submission copied to clipboard
qca-dataset-submission copied to clipboard
Missing index mapping QCArchive dataset names to datasets in this repo
The folks in the lab have been trying to make use of datasets in QCArchive, but it appears to be impossible to track down the provenance information for some of those datasets.
For example, right now, I'm trying to identify which dataset corresponds to the QCArchive dataset named OpenFF Optimization Set 1:
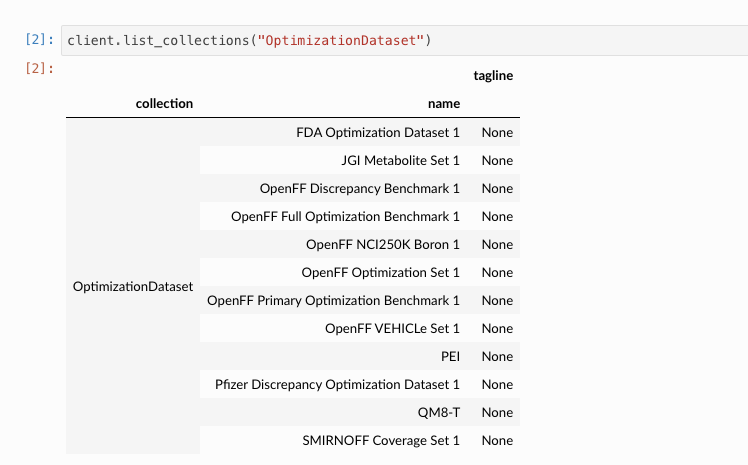 However, that text does not appear at all in this repo.
However, that text does not appear at all in this repo.
Is there a master mapping we could use between dataset names in QCArchive and the directories containing input files used to generate them and associate discussions here in this repo?
Thanks for relaying this to us @jchodera. We weren't aware of this pain point, and speaking for myself I tend to do a grep -r "<dataset_name>" to get back which submission directory(s) corresponds to the dataset I'm interested in understanding more about. That may help users in the short term.
We can do better than this, though. I think having a file featuring a table giving columns "dataset name" and "submission directory" would be sufficient, and we can make adding an entry to this part of our PR checklist for new submissions (similar to a changelog). We could call it INDEX.md, unless someone has a better name.
@trevorgokey, @jthorton, do you agree with this approach?
I am pretty sure these are panda DataFrame woes; the dataset is there but, by default, prints a truncated list when you work with it the "right way" using the Index objects.
I read somewhere the following snippet is bad practice for going over large datasets, but it (called list.py in some bin dir) has helped me a bunch:
import qcfractal.interface as ptl
def main():
client = ptl.FractalClient()
for row in client.list_collections().iterrows():
print(row[0][0], row[0][1])
if __name__ == "__main__":
main()
One could could alternatively set pd.options.display.max_rows
Yes thanks for pointing this out, this is always going to be a problem with the older datasets however new datasets have the long description address saved into the metadata. @dotsdl I think this is a good idea however looking into this one I think we may have accidentally lost the input molecules for this dataset during the reorganization, I think the scipts and inputs should be in this folder but it looks like only the hessian script was kept. I might have missed it in another place however.
I am pretty sure these are panda DataFrame woes; the dataset is there but, by default, prints a truncated list when you work with it the "right way" using the Index objects.
It's the other way around, @trevorgokey : The QCArchive dataframe contains a dataset called OpenFF Optimization Set 1, but I have no idea which of the directories in this repository contains the scripts and input files and documentation associated with this dataset.
@dotsdl : Thanks for the grep idea! I agree that having a table would be super useful.
@jthorton : Would be great if we could recover the input files if at all possible!