Numba function works randomly with same given input, is this a bug?
I checked and there is no other issue resembling mine.
Reporting a bug
- [x ] I have tried using the latest released version of Numba (most recent is visible in the change log (https://github.com/numba/numba/blob/main/CHANGE_LOG).
- [x ] I have included a self contained code sample to reproduce the problem. i.e. it's possible to run as 'python bug.py'.
Stackoverflow issue: Numba function works randomly with same given input, is this a bug?
Description
I wrote a function called not_test in Numba to take a list of 2d arrays that is a drainage network, then I get an imaginary water drop to each point for routing from the figure below. The point of the code is to get the path of the drop for every possible drainage stream.
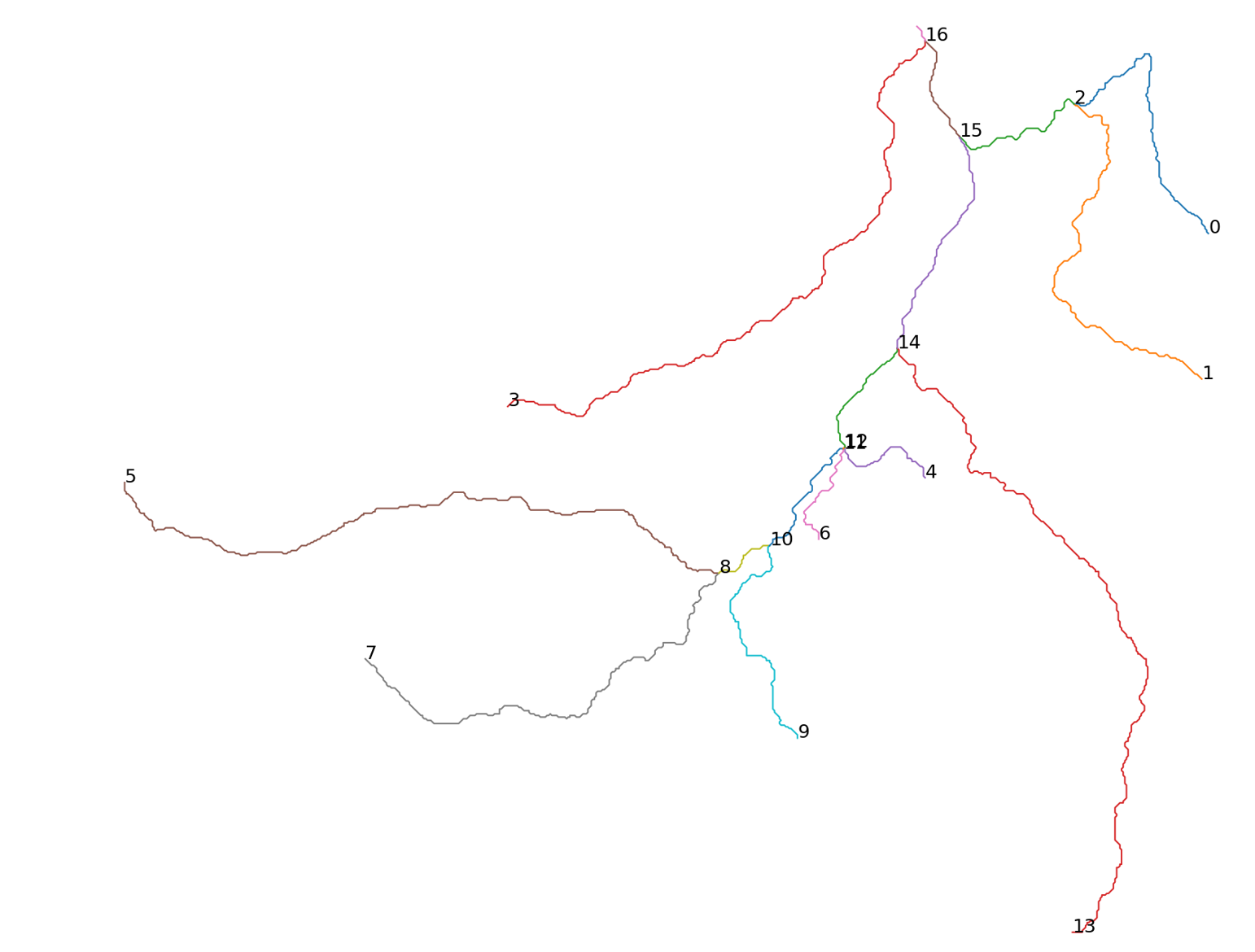
Expected Results
These are the results I am getting, the integer sequence is the routing stream a water drop would take if it falls in the start of the stream, eg. falls in point 1 then, routing stream would be [16, 15, 2, 1].
[[16, 3],
[16, 15, 2, 0],
[16, 15, 2, 1],
[16, 15, 14, 13],
[16, 15, 14, 12, 4],
[16, 15, 14, 12, 11, 6],
[16, 15, 14, 12, 11, 10, 9],
[16, 15, 14, 12, 11, 10, 8, 5],
[16, 15, 14, 12, 11, 10, 8, 7]]
Problem
The code works on normal python and it works as well when it is compile with Numba. The problem comes if you ran several times the code compile with Numba, some times this gives an error and sometimes it does work. (see error in figure below)
I have not been able to debug the code in Numba and it gives no error in python mode. And it does not show any particular error in the python console or pycharm run, and there is no error with boundscheck=True. debug=True. It just stops.
The code that is commented is sure not to be part of the issue I am experiencing.
I would really like to been able to use Numba on this function because it has a 653X speed up, and this function will ran around 5k times, this would mean:
with Numba: 0.0015003681182861328s per run -> 7.5s total time
with Python: 0.9321613311767578s per run -> 1.3 hours total time
Using Numba is a BIG help in this particular issue, so I would appreciate any help, because normal python would not work for the application usage.
Error exmaple
The error does not give an exception error and it gives a arbitrary place to stop the code eg. [6 10] or [8 9]
in Pycharm error:
Now
0.0
0.2295396327972412
[16]
[ 3 15]
[ 3 15]
[ 2 14]
[0 1]
[12 13]
Process finished with exit code -1073740940 (0xC0000374)
in Pycharm no error:
Now
0.0
0.2430422306060791
[16]
[ 3 15]
[ 3 15]
[ 2 14]
[0 1]
[12 13]
[ 4 11]
[ 4 11]
[ 4 11]
[ 6 10]
[ 6 10]
[8 9]
[5 7]
[[16, 3], [16, 15, 2, 0], [16, 15, 2, 1], [16, 15, 14, 13], [16, 15, 14, 12, 4], [16, 15, 14, 12, 11, 6], [16, 15, 14, 12, 11, 10, 9], [16, 15, 14, 12, 11, 10, 8, 5], [16, 15, 14, 12, 11, 10, 8, 7]]
0.0016527080535889
Process finished with exit code 0
Code
link to file: test.npy
import numpy as np
#from pypiper import RUT_5
import numba
def convert2(x, dtype=np.float64):
try:
# Try and convert x to a Numpy array. If this succeeds
# then we have reached the end of the nesting-depth.
y = np.asarray(x, dtype=dtype)
except:
# If the conversion to a Numpy array fails, then it can
# be because not all elements of x can be converted to
# the given dtype. There is currently no way to distinguish
# if this is because x is a nested list, or just a list
# of simple elements with incompatible types.
# Recursively call this function on all elements of x.
y = [convert2(x_, dtype=dtype) for x_ in x]
# Convert Python list to Numba list.
y = numba.typed.List(y)
return y
@numba.njit('(ListType(float64[:, ::1]), float64[:])')
def not_test(branches, outlet):
# get len of branches
_len_branches = len(branches)
# # empty array
# d_array = np.empty(shape=_len_branches, dtype=np.float64)
# # set outlet coordinates as arrays
# x_outlet, y_outlet = outlet
# x_outlet, y_outlet = np.array([x_outlet]), np.array([y_outlet])
#
# # get min distance from branches
# for pos in numba.prange(_len_branches):
# # get current branch
# branch = branches[pos]
# # get min distance from outlet point
# d_min = RUT_5.nb_cdist(branch, x_outlet, y_outlet).min()
# # add to array
# d_array[pos] = d_min
#
# #get index for minimun distance
# index_branch = np.argmin(d_array)
index_branch = 16
#remove initial branch
update_branches = branches.copy()
del update_branches[index_branch]
#define arrays
not_read = np.empty(shape=0, dtype=np.int64)
paths_update = [[np.int(x)] for x in range(0)]
points = np.empty(shape=(2, 2))
a_list = [np.int(x) for x in range(0)]
# avoid from loop
not_read = np.append(index_branch, not_read)
# iterable in loop
iterable = not_read.copy()
# conditions
cond = 0
cont = 0
while cond == 0:
for pos_idx in iterable:
print(iterable)
if cont > 0:
paths = paths_update.copy()
branch = branches[pos_idx]
points[0] = branch[0]
points[1] = branch[-1]
for point in points:
for pos_j in range(_len_branches):
if pos_j not in not_read:
diff = np.sum(point - branches[pos_j], axis=1)
if 0 in diff:
a_list.append(pos_j)
if cont == 0:
paths = [[pos_idx] + [i] for i in a_list]
paths_update = paths.copy()
cont = cont + 1
not_read = np.append(not_read, a_list)
iterable = np.array(a_list)
a_list = [np.int(x) for x in range(0)]
else:
if len(a_list):
path_arr = [_i for _i in paths if pos_idx in _i]
for path in path_arr:
for conexion in a_list:
temp_list = path.copy()
temp_list.append(conexion)
paths_update.append(temp_list)
paths_update.remove(path)
not_read = np.append(not_read, a_list)
iterable = np.array(a_list)
a_list = [np.int(x) for x in range(0)]
else:
continue
if len(branches) == len(np.unique(not_read)):
cond = 1
return paths
if __name__ == '__main__':
print('Now')
branches = np.load('test.npy', allow_pickle=True).item()
x_snap, y_snap = 717110.7843995667, 9669749.115011858
import time
t0 = time.time()
arr = []
for pos, branch in enumerate(branches.features):
arr.append(list(branch.geometry.coordinates))
print(time.time() - t0)
t0 = time.time()
arr = convert2(arr)
print(time.time() - t0)
t0 = time.time()
outlet = np.array([x_snap, y_snap])
print(not_test(branches=arr, outlet=outlet))
print(time.time() - t0)
@CHELOBV thank you for submitting this. I have started to triage this issue. I downloaded the file test.npy, however, when running the code I receive the error:
💣 zsh» python issue_7890.py
Now
Traceback (most recent call last):
File "/Users/esc/git/numba/issue_7890.py", line 120, in <module>
branches = np.load('test.npy', allow_pickle=True).item()
File "/Users/esc/miniconda3/envs/numba-dev/lib/python3.9/site-packages/numpy/lib/npyio.py", line 440, in load
return format.read_array(fid, allow_pickle=allow_pickle,
File "/Users/esc/miniconda3/envs/numba-dev/lib/python3.9/site-packages/numpy/lib/format.py", line 748, in read_array
array = pickle.load(fp, **pickle_kwargs)
ModuleNotFoundError: No module named 'geojson'
python issue_7890.py 5.57s user 0.34s system 78% cpu 7.529 total
I presume that a pip install geojson will solve it.
After installing geojson I ran the supplied reproducer and data:
💣 zsh» python issue_7890.py
Now
3.314018249511719e-05
0.21585464477539062
[16]
[ 3 15]
[ 3 15]
[ 2 14]
[0 1]
[12 13]
[ 4 11]
[ 4 11]
[ 4 11]
[ 6 10]
[ 6 10]
[8 9]
[5 7]
python(65338,0x10ba0bdc0) malloc: Incorrect checksum for freed object 0x7fe893c0ac80: probably modified after being freed.
Corrupt value: 0x5fffffffffffffff
python(65338,0x10ba0bdc0) malloc: *** set a breakpoint in malloc_error_break to debug
Also, this came back with:
💣 zsh» echo $?
134
``
Which would indicate that program received a `SIGABRT`.
After running this with boundschecking=True I received yet another error message:
💣 zsh» python issue_7890.py
Now
3.409385681152344e-05
0.18205642700195312
[16]
[ 3 15]
[ 3 15]
[ 2 14]
[0 1]
[12 13]
[ 4 11]
[ 4 11]
[ 4 11]
[ 6 10]
[ 6 10]
[8 9]
[5 7]
SystemError: /opt/concourse/worker/volumes/live/c1a1a6ef-e724-4ad9-52a7-d6d68451dacb/volume/python-split_1631807121927/work/Objects/listobject.c:138: bad argument to internal function
The above exception was the direct cause of the following exception:
Traceback (most recent call last):
File "/Users/esc/git/numba/issue_7890.py", line 136, in <module>
print(not_test(branches=arr, outlet=outlet))
SystemError: CPUDispatcher(<function not_test at 0x7fb9eea621f0>) returned a result with an error set
python(66602,0x111ba2dc0) malloc: tiny_free_list_remove_ptr: Internal invariant broken (prev ptr of next): ptr=0x7fb9f0d76890, next_prev=0x7fb9f0d76880
python(66602,0x111ba2dc0) malloc: *** set a breakpoint in malloc_error_break to debug
[1] 66602 abort (core dumped) python issue_7890.py
python issue_7890.py 5.08s user 4.15s system 73% cpu 12.614 total
I debugged this some more, and found that the array may in fact be an object array. I don't think that Numba supports object-arrays very well.
In [3]: numpy.load("test.npy", allow_pickle=False)
---------------------------------------------------------------------------
ValueError Traceback (most recent call last)
<ipython-input-3-30c0219c8f76> in <module>
----> 1 numpy.load("test.npy", allow_pickle=False)
~/miniconda3/envs/numba-dev/lib/python3.9/site-packages/numpy/lib/npyio.py in load(file, mmap_mode, allow_pickle, fix_imports, encoding)
438 return format.open_memmap(file, mode=mmap_mode)
439 else:
--> 440 return format.read_array(fid, allow_pickle=allow_pickle,
441 pickle_kwargs=pickle_kwargs)
442 else:
~/miniconda3/envs/numba-dev/lib/python3.9/site-packages/numpy/lib/format.py in read_array(fp, allow_pickle, pickle_kwargs)
741 # The array contained Python objects. We need to unpickle the data.
742 if not allow_pickle:
--> 743 raise ValueError("Object arrays cannot be loaded when "
744 "allow_pickle=False")
745 if pickle_kwargs is None:
ValueError: Object arrays cannot be loaded when allow_pickle=False
@CHELOBV you mentioned, that this does happen sometimes only? For me, the code and array crash reproducibly every time.
So far I was on 3.9 on OSX Catalina on x86_64, now, switching to 3.10 on Linux x86_64, I get:
💥 zsh» python issue_7890.py
Now
4.206834316253662
dog
[16]
[ 3 15]
[ 3 15]
[ 2 14]
[0 1]
[12 13]
[1] 4424 segmentation fault (core dumped) python issue_7890.py
python issue_7890.py 6,08s user 0,12s system 61% cpu 10,041 total
Which is a slightly different error.
So, I don't really understand the code very well, unfortunately, but I did manage to eliminate the segfault:
By commenting out the line paths_update.remove(path) on line 102, I can get the script to run through.
The results with Numba:
💥 zsh» python issue_7890.py
Now
4.00543212890625e-05
0.19878649711608887
[16]
[ 3 15]
[ 3 15]
[ 2 14]
[0 1]
[12 13]
[ 4 11]
[ 4 11]
[ 4 11]
[ 6 10]
[ 6 10]
[8 9]
[5 7]
[[16, 3], [16, 15], [16, 15, 2], [16, 15, 14], [16, 15, 2, 0], [16, 15, 2, 1], [16, 15, 14, 12], [16, 15, 14, 13], [16, 15, 14, 12, 4], [16, 15, 14, 12, 11], [16, 15, 14, 12, 11, 6], [16, 15, 14, 12, 11, 10], [16, 15, 14, 12, 11, 10, 8], [16, 15, 14, 12, 11, 10, 9], [16, 15, 14, 12, 11, 10, 8, 5], [16, 15, 14, 12, 11, 10, 8, 7]]
0.0013725757598876953
And without Numba, with the @njit decorator commented out:
💥 zsh» python issue_7890.py
Now
5.507469177246094e-05
0.3475668430328369
[16]
[ 3 15]
[ 3 15]
[ 2 14]
[0 1]
[12 13]
[ 4 11]
[ 4 11]
[ 4 11]
[ 6 10]
[ 6 10]
[8 9]
[5 7]
[[16, 3], [16, 15], [16, 15, 2], [16, 15, 14], [16, 15, 2, 0], [16, 15, 2, 1], [16, 15, 14, 12], [16, 15, 14, 13], [16, 15, 14, 12, 4], [16, 15, 14, 12, 11], [16, 15, 14, 12, 11, 6], [16, 15, 14, 12, 11, 10], [16, 15, 14, 12, 11, 10, 8], [16, 15, 14, 12, 11, 10, 9], [16, 15, 14, 12, 11, 10, 8, 5], [16, 15, 14, 12, 11, 10, 8, 7]]
0.5873363018035889
My guess was that something was being removed twice or from a location where it should not be removed. I looked into the code some more and saw that paths_update isn't actually being used for anything at all. So probably all statements related to this variable can be removed. In fact, I think there are quite a few opportunities for removing redundant code from this example.
@CHELOBV while this may be good news for you, it still means there could be a latent bug in the Numba. So I think it would make sense to reduce the code to trigger the issue to an absolute minimum. This is called "crafting a reproducer". This would give us additional hints as to what is going on and if this is indeed a Numba bug.
I ran this again with the line paths_update.remove(path) intact and under pure Python, and I get a different result:
💣 zsh» python issue_7890.py
Now
4.00543212890625e-05
0.3786599636077881
dog
[16]
[ 3 15]
[ 3 15]
[ 2 14]
[0 1]
[12 13]
[ 4 11]
[ 4 11]
[ 4 11]
[ 6 10]
[ 6 10]
[8 9]
[5 7]
[[16, 3], [16, 15, 2, 0], [16, 15, 2, 1], [16, 15, 14, 13], [16, 15, 14, 12, 4], [16, 15, 14, 12, 11, 6], [16, 15, 14, 12, 11, 10, 9], [16, 15, 14, 12, 11, 10, 8, 5], [16, 15, 14, 12, 11, 10, 8, 7]]
0.4874241352081299
Probably this means, that the statement does have an effect, contrary to my previous statement.
Unfortunately, I am out of time for now. If someone would like to pickup debugging this, it probably has something to do with the line paths_update.remove().
Hi, thank you for reviewing this. The code runs randomly for me with the @njit decorator. I am running this on windows 10 and python 3.9.7 and my numba is 0.54.1. This behavior does not occur in pure python, but it could be something I am doing wrong that is not allowed when working with numba.
The path_update.remove(path) remove unwanted paths for the streams, so I you compare against my desire result you will see that this [16, 15], [16, 15, 2], [16, 15, 14] (among others ) should not be in the code. t does have and effect as it avoid changing the list its currently lopping.
Thanks for the effort. Marcelo.
Hi, I changed the code as below. Now it works just fine with the @njit decorator, it is not ideal as it iterates over a set of paths that should not be there but it get the jod done and is quite faster than normal python.
The error definitely is with the list.remove(value) and the list.pop(index), as I tried both with the same results as the initially proposed with the remove method and a total error with the pop method. They both work if these methods are avoided.
notice the line del update_branches[index_branch] works.
@numba.njit('(ListType(float64[:, ::1]), float64[:])')
def not_test(branches, outlet):
# get len of branches
_len_branches = len(branches)
# # empty array
# d_array = np.empty(shape=_len_branches, dtype=np.float64)
# # set outlet coordinates as arrays
# x_outlet, y_outlet = outlet
# x_outlet, y_outlet = np.array([x_outlet]), np.array([y_outlet])
#
# # get min distance from branches
# for pos in numba.prange(_len_branches):
# # get current branch
# branch = branches[pos]
# # get min distance from outlet point
# d_min = RUT_5.nb_cdist(branch, x_outlet, y_outlet).min()
# # add to array
# d_array[pos] = d_min
#
# #get index for minimun distance
# index_branch = np.argmin(d_array)
index_branch = 16
#remove initial branch
update_branches = branches.copy()
del update_branches[index_branch]
#define arrays
not_read = np.empty(shape=0, dtype=np.int64)
paths_update = [[np.int(_)] for _ in range(0)]
paths_remove = [np.int(_) for _ in range(0)]
points = np.empty(shape=(2, 2))
a_list = [np.int(x) for x in range(0)]
diff = np.empty(shape=0, dtype=np.int64)
# avoid from loop
not_read = np.append(index_branch, not_read)
# iterable in loop
iterable = not_read.copy()
# conditions
cond = 0
cont = 0
while cond == 0:
for pos_idx in iterable:
if cont > 0:
paths = paths_update.copy()
branch = branches[pos_idx]
points[0] = branch[0]
points[1] = branch[-1]
for point in points:
for pos_j in numba.prange(_len_branches):
if pos_j not in not_read:
diff = np.sum(point - branches[pos_j], axis=1)
if len(diff[diff == 0]) > 0:
a_list.append(pos_j)
if cont == 0:
paths = [[pos_idx] + [i] for i in a_list]
paths_update = paths.copy()
cont = 1
not_read = np.append(not_read, a_list)
iterable = np.array(a_list)
a_list = [np.int(x) for x in range(0)]
else:
if len(a_list):
for pos, path in enumerate(paths):
if pos_idx in path:
for conexion in a_list:
temp_list = path.copy()
temp_list.append(conexion)
paths_update.append(temp_list)
paths_remove.append(pos)
not_read = np.append(not_read, a_list)
iterable = np.array(a_list)
a_list = [np.int(x) for x in range(0)]
if len(branches) == len(np.unique(not_read)):
cond = 1
paths = [_ for _i, _ in enumerate(paths) if _i not in paths_remove]
return paths
@CHELOBV thank you for updating this issue. I still don't understand why exactly the remove call causes the issue here. My assumption is that it has something to do with the nested lists and how they are copied.
I ran it again today on my mac and received a slightly different error message:
Traceback (most recent call last):
File "/Users/vhaenel/git/numba/issue_7890.py", line 136, in <module>
print(not_test(branches=arr, outlet=outlet))
SystemError: CPUDispatcher(<function not_test at 0x7fde04d621f0>) returned a result with an error set
python(21919,0x11aafedc0) malloc: tiny_free_list_remove_ptr: Internal invariant broken (prev ptr of next): ptr=0x7fde07e1eaa0, next_prev=0x7fde07e1ea90
python(21919,0x11aafedc0) malloc: *** set a breakpoint in malloc_error_break to debug
With a hint of:
tiny_free_list_remove_ptr
@CHELOBV I am glad you managed to find a solution that is better than pure-python already. If anyone would like to continue to debug this, I would recommend trying to craft a minimal reproducer around the remove call to work out why that failed.
I could try to provide this, if it helps at all. greetings, Chelo.
@CHELOBV that would be totally awesome. If you have spare time and inclination to look into acrafting a reproducer that would help the dev team a lot as there may really be a Numba bug at fault here. Our good friend Mathew Rocklin wrote the following blog post on how to craft a minimal reproducer, which is quite helpful:
https://matthewrocklin.com/blog/work/2018/02/28/minimal-bug-reports
Good luck, if you do decide to take this on!
This issue is marked as stale as it has had no activity in the past 30 days. Please close this issue if no further response or action is needed. Otherwise, please respond with any updates and confirm that this issue still needs to be addressed.
@esc @CHELOBV what's the status of this issue? Is there still a bug to report or can it be closed as resolved?
@stuartarchibald I just tried to run the reproducer and it still fails on current main it'll need a minimum reproducer to be extracted. I've added this to my queue for now.
This issue is marked as stale as it has had no activity in the past 30 days. Please close this issue if no further response or action is needed. Otherwise, please respond with any updates and confirm that this issue still needs to be addressed.
Hi, I missed the message from github, this is still an issue, @esc what do you need ? because the exmaple of the issue is self contained. Sorry if I dont understand the problem here. Grettings, Marcelo.
Hi, I missed the message from github, this is still an issue, @esc what do you need ? because the exmaple of the issue is self contained. Sorry if I dont understand the problem here. Grettings, Marcelo.
I would need a minimum working reproducer. The example is currently still too large to debug effectively. See also: https://matthewrocklin.com/blog/work/2018/02/28/minimal-bug-reports
Hi @esc , do you mean the size of the data ? because the actual code is a little as posible, I read the doc you send me, the thing says no to is to attach a file as I did. Grettings, Marcelo.
Hi @esc , do you mean the size of the data ? because the actual code is a little as posible, I read the doc you send me, the thing says no to is to attach a file as I did. Grettings, Marcelo.
OK, thank you for updating! I was under the impression that the reproducer may be reducible in size to eliminate more potential sources for the issue.
This issue is marked as stale as it has had no activity in the past 30 days. Please close this issue if no further response or action is needed. Otherwise, please respond with any updates and confirm that this issue still needs to be addressed.
This issue is marked as stale as it has had no activity in the past 30 days. Please close this issue if no further response or action is needed. Otherwise, please respond with any updates and confirm that this issue still needs to be addressed.
I will remove the stale label again. I still hope to find time to debug this, someday.