ENH: List instead of merely counting private mutations
Nextclade lets users know how many private mutations are in a sequence but it would be great if it could generate a list of these private mutations. This could be either flagging them in the list of changes with a * or (P), or listing them separately a new column of the csv report.
Why is this useful? Number of private mutations and mutation clusters are both really helpful QC metrics that tell us about sequence quality and flag potential errors. Private mutations and clusters can sometimes be due to issues with trimming or quality issues at the ends of reads combined with low coverage at these sites leading to them being called in consensus. Having a list of where these private mutations (and maybe also mutation clusters) are will help us to go back to the bams or vcfs, review read support for each mutation, determine whether they are real and clean up the sequence.
Thanks, Pavitra! I like this suggestion and it should be pretty straightforward to implement.
Hey Richard! Sorry to jump in, but is this feature implemented in the web version? It would be great being able to get the list of private mutations.
Thanks for a super helpful tool!
Gonzalo
@GonzaloYebra !!!!!! 😁 It is lovely to see you, even if it's on a github issue thread! 🙃
Haha likewise Emma! Any little corner would do in this day and age :P
@proychou We've implemented this for a while now - both in web as well as in the TSV export
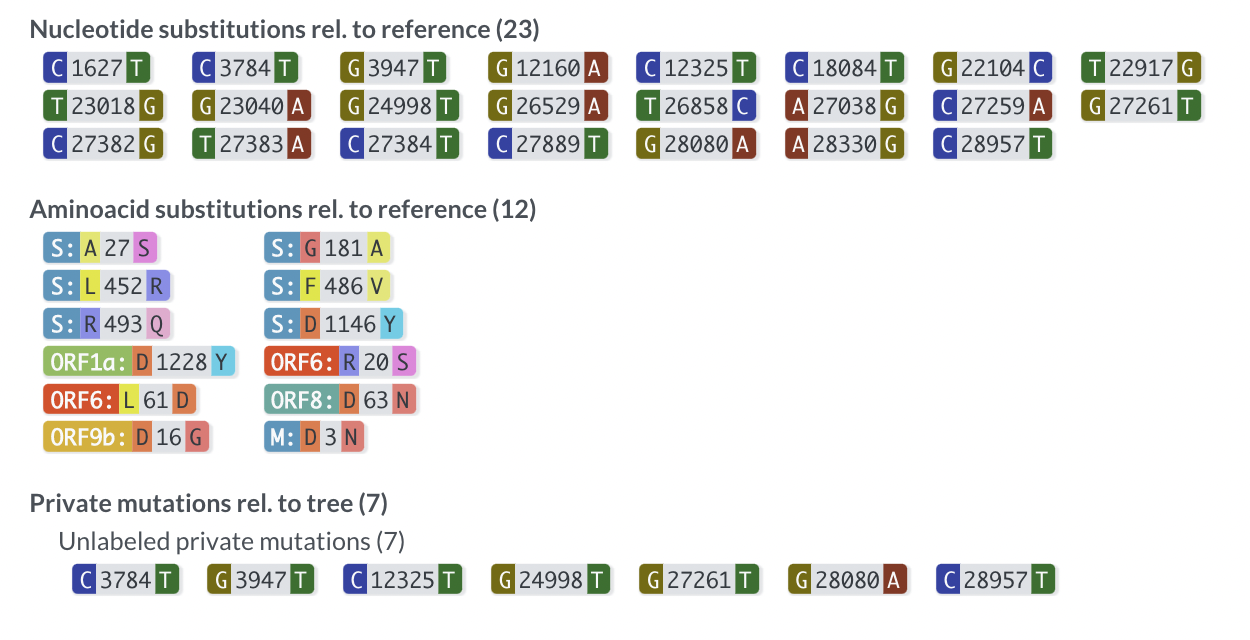

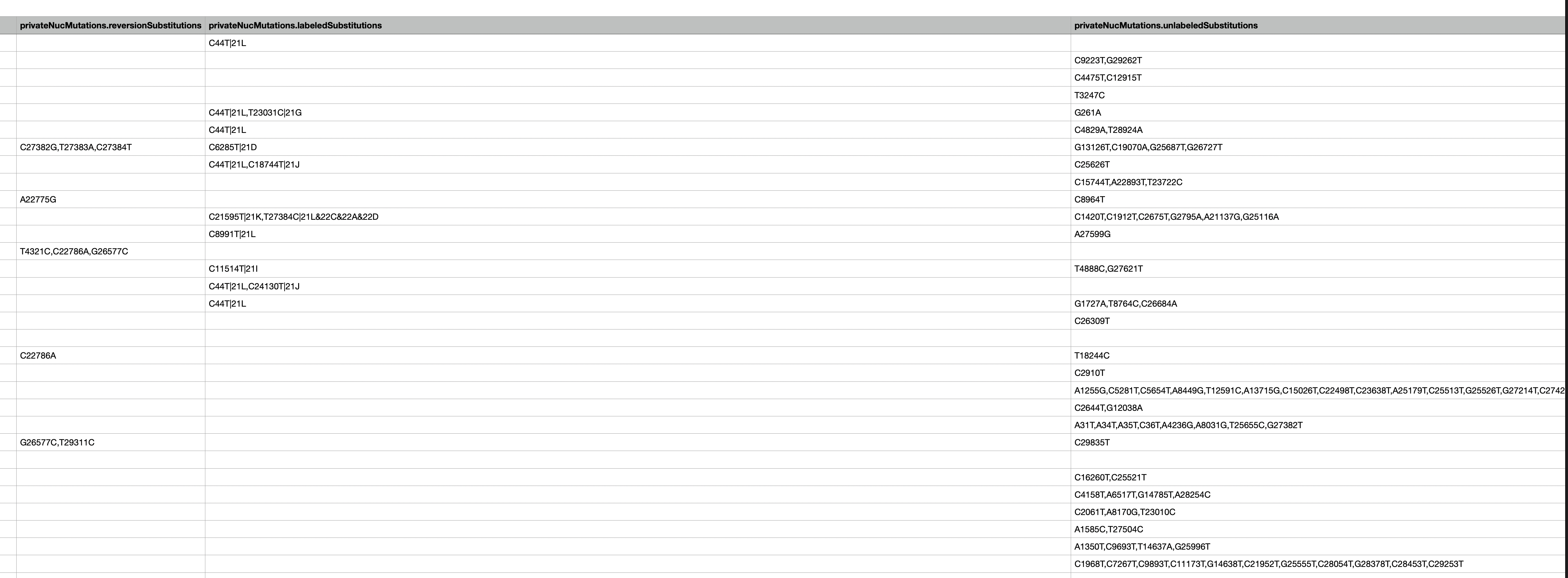
Thanks for the idea!