Built-in, standard, and template montages, and layouts 🤯
Hello,
now this is something that has been confusing me for quite a while, and I'm quite certain it's something that will bite new EEG users who join the MNE-Python side of things (for we have cookies! 🍪):
When trying to learn more about EEG channel layouts (ha! I said it!), one might actually look for that term – layout. But in MNE, it has a different meaning than EEG users would expect. This is the reason why I felt urged to add a Note box to the sensor locations tutorial, stating that if you're working with EEG recordings, you should only be interested in montages and not layouts:
If you’re working with EEG data exclusively, you’ll want to use Montages, not layouts. Idealized montages (e.g., those provided by the manufacturer, or the ones shipping with MNE-Python mentioned below) are typically referred to as template montages.
Ok, now with that out of the way, let's look at montages.
We ship a number of built-in montages, whose names can be retrieved via a function named get_builtin_montages() – although the name of that function seems to suggest it will return an actual montage, not just a montage name. Oh well, legacy stuff, backward-compat, let's just keep on moving.
Sooo… now that we've picked our favorite, we can pass that montage name to a function called … make_standard_montage(). So suddenly we go from builtin_montage to standard_montage. scratches head
Oh well, the developers probably know what they're doing, right? Let's keep on moving!
Being a standard-conforming user of EEG, my electrodes are arranged according to the 10-20 or 10-05 system. Neat!, I think, MNE ships with a standard_1020 builtin montage! So let's use that one and visualize it:
# %%
import mne
m = mne.channels.make_standard_montage('standard_1020')
m.plot(show_names=True)
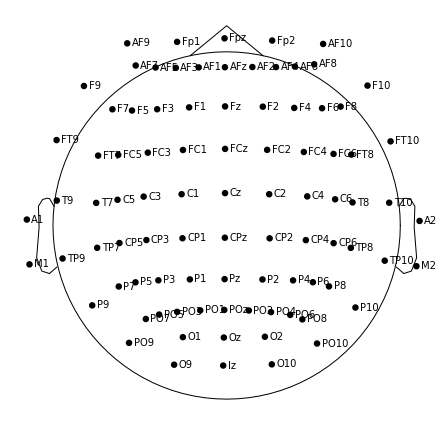
Ok, looks … weird, but after studying the tutorial and API docs, I realize there's this sphere parameter I should probably use.
m.plot(show_names=True, sphere='eeglab')
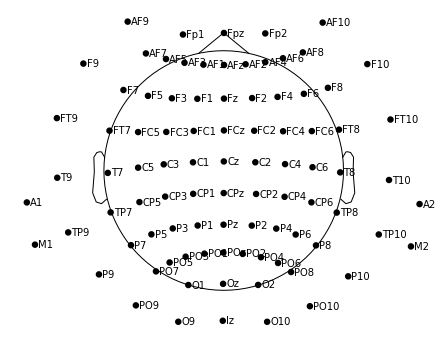
… wait, what? This still looks unexpected – why's aren't the electrodes between T7 and T8 aligned in a straight horizontal line??
Again, I start checking the docs … and the only hint I find is hidden in the make_standard_montage() docstring under the description of a parameter I didn't even use (and will never even find out about if I pass the montage name to set_montage() directly):
head_size : float | None | str The head size (radius, in meters) to use for spherical montages. Can be None to not scale the read sizes. ``'auto'`` (default) will use 95mm for all montages except the ``'standard*'``, ``'mgh*'``, and ``'artinis*'``, which are already in fsaverage's MRI coordinates (same as MNI).
So apparently those standard_* montages are so overly standard that they're special.
Since I'm new to MNE, I don't even have the slightest clue what this fsaverage is and why its "MRI coordinates" have anything to do with how sensors are aligned on what I thought would be a standard montage.
So I use one of those easycap or biosemi montages because I've heard of this before, and now things start to be looking good:
m = mne.channels.make_standard_montage('easycap-M1')
m.plot(show_names=True, sphere='eeglab')
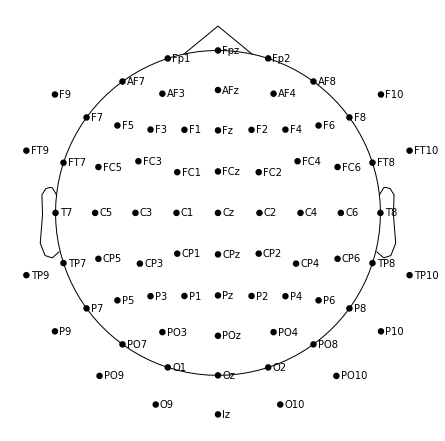
From studying the MNE-BIDS docs I know of the term "template montage", and the docs establish a direct link between this term and the "standard montages" in MNE-Python (note that the term already occurred in the Note box I mentioned at the very beginning; however I only added this box … yesterday)
The *electrodes.tsv and *coordsystem.json files in BIDS are intended to carry information about digitized (i.e., measured) electrode positions on the scalp of the research subject. Do not (!) use these files to store “template” or “idealized” electrode positions, like those that can be obtained from mne.channels.make_standard_montage()!
And now my mind is fully blown:
- "template montages" are standard montages
- "standard montages" are "built-in" montages
- the "standard montages" whose name starts with the literal term
standardare morphed into a super odd shape by fitting them onto thisfsaverageMRI that I've never heard of before, making them anything but "standard" in comparison to the "templates" I've seen in the past being provided by cap manufacturers
And I'm just like: EHLP ! 😱😱
Isn't there anything we can do to salvage this mess?
cc @sappelhoff @cbrnr
Reading the EEGLAB wiki, I'm starting to think that one of the things we should do is recommend users apply one of the standard (or other MNI-morphed) montages if their electrode setup was according to the 10-20 or 10-05 system and they're planning to perform source estimation and they don't have individual MRIs – regardless of the actual cap / EEG system they used in the lab:
As of 2021, the default channel location file for electrode position is the MNI file, which is best suited for source localization. Before 2021, it was the BESA spherical location file.
https://eeglab.org/tutorials/04_Import/Channel_Locations.html
If all your electrode locations are within the International 10-20 System (which is the case here), you may use the standard channel coordinates associated with the head model. [...] Select Use MNI coordinate file for the BEM DIPFIT model.
https://eeglab.org/tutorials/09_source/Model_Settings.html
I have to say I still don't understand why the montage plot doesn't show T7<->Cz<->T8 in a straight line, but that's probably because I don't fully understand how that fit was achieved. Did one start out with a sphere with optimally placed electrodes according to 10-05 / 10-20, and then moved the electrodes radially until they were all placed on the scalp level of fsaverage? Could anybody please explain?
Does this also mean that we must actually advice users not to use any non-MNI-morphed built-in montages if they're planning to do source estimation?
(I know that @agramfort strongly believes one shouldn't do inverse modeling on EEG without measured sensor locations and and individual MRs scan anyway, but people do do that and it actually yields reasonable results, depending on the expectations one sets a-priori :))
Alright, it seems like there are several issues relating to montages, so I'll comment just here.
I agree that the layout/montage distinction is confusing. Layouts are really montages in 2D, so potentially we could unify both as montages with a 3D or 2D value in some attribute.
The main point of confusion for me is how we ship our montages. It is not intuitive that standard_1005 and standard_1020 are not template montages (i.e. computed on a sphere). I think that we should include a utility function which creates template montages from given channel labels – exactly what @sappelhoff has implemented a long time ago (https://github.com/sappelhoff/eeg_positions). We've discussed this before but never reached a decision.
My suggestion is to remove all montage files and create (compute) them from scratch. That way, we know for sure that we're dealing with template montages. We only need to include montages that we cannot derive computationally, i.e. a 10–05 montage on the MNI averaged head. I would also make sure to carefully distinguish between
- template montages (computed on a sphere),
- realistic montages (computed e.g. on fsaverage),
- and individual montages (which we cannot include of course).
My suggestion is to remove all montage files and create (compute) them from scratch.
But then we run into issues regarding channel naming, no? Because different caps sometimes have different names for their sensors, even if they're placed in the exact same location … so we'd still need to ship cap-specific channel names. Or am I mistaken?
Aside from this, I very much love the idea!
Layouts are really montages in 2D, so potentially we could unify both as montages with a 3D or 2D value in some attribute.
Oh boy, this never really occurred to me. So the only reason we're even shipping "layouts" is to avoid the trouble with 3D->2D projection that can never make everybody happy?
We can have duplicate locations with different names (we're doing this already I think). So our 10–05 template montage just needs to include all possible labels that we know of? But I'd also be OK with cap-specific montages. After all, 10–05 names are already defined. If some specific cap uses different naming conventions, we can ship a separate montage (although in the background we're really using the same locations).
Oh boy, this never really occurred to me. So the only reason we're even shipping "layouts" is to avoid the trouble with 3D->2D projection that can never make everybody happy?
That's my understanding. But I might be wrong, which would even further support your point that the current situation is confusing 😄. In some layouts, we want to optimize for space and not for correct projection, but we could do this with special montages too.
We could also de-emphasize layouts, because I have never needed to use a layout before. Montages are way more important for EEG analysis, and layouts could be considered a specialized class which is nice for generating 2D plots.
We could also de-emphasize layouts, because I have never needed to use a layout before.
But I suppose that's just because you're working with EEG data. My understanding up until 30 mins ago was that layouts are (only) meant for 2D viz of MEG sensors. But I just saw that there's also a Biosemi layout, and now I just want to cry. 😭
Yes, maybe so, I've been ignoring all MEG stuff so far (sorry folks 😆). And yes, there are layouts for EEG (which I never needed). So we should find out why layouts are needed for MEG, and if they could be replaced by montages.
I agree that the layout/montage distinction is confusing. Layouts are really montages in 2D, so potentially we could unify both as montages with a 3D or 2D value in some attribute.
At some point we decided something along the lines of:
-
plot_topomap(and any 2D plotting with head + sensors) should use a standardized mapping between the 3D sensor locations and a flattened 2D view (controlled by stuff likesphere) - A layout is an arbitrary mapping between channel names and axes in a 2D space
- A DigMontage (and "montage" in general) is a set of 3D positions plus channel names (more or less)
I'd rather not revisit / redefine these terms if possible. If someone is really motivated to change these, can you please search through all previous discussions, summarize them, and link to them here? Until someone does this leg work, I'd prefer not to discuss this further here. There might very well be things that we should change here, but we risk revisiting previously raised points if nobody goes through this effort.
And now my mind is fully blown: ... "template montages" are standard montages ...
Maybe a way to start here is to
- Add entries to
glossarythat we can all agree on / reflect what the current state of affairs is meant to be - Make PRs to fix our language to use and link to these terms as much as possible
So we should find out why layouts are needed for MEG, and if they could be replaced by montages.
As per the def above, Layouts allow you to divorce the physical position of the sensors from how they are shown. Sometimes this is absolutely necessary, like for Neuromag MEG systems where sensor triplets (planar1, planar2, mag) have the same physical loc. Other times it's just convenient for other reasons, e.g., a nicely idealized/separated one, if our systematic flattening does not provide it -- I assume for example this was the motivation for https://github.com/mne-tools/mne-python/pull/10627
Thanks for clarifying the layout vs. montage distinction – this should go into the glossary (in exactly that level of detail with the MEG example). No need to redefine then.
@larsoner what do you think about the template (computed) vs. realistic (defined on averaged data) vs. individual montages? Besides clarifying terminology, I think our built-in montages should really be calculated from scratch (instead of read from a file). I usually have two applications for montages:
- Make pretty visualizations (for which I want to use template montages computed with a sphere model)
- Source localization (for which I would either use individually measured montages or a montage based on fsaverage, but even a spherical montage would work OK if I don't have subject-specific MRIs).
Currently, there is no way to tell in which category a given montage belongs to (e.g. standard_1020 is a measured one I think, but it sounds like it should be a computed one – but we could have both of course).
I think our built-in montages should really be calculated from scratch
I don't think we have a way to compute the MNI/realistic locations from scratch, right? So this seems like a two-part argument: 1) our positions should default to idealized/spherical, and 2) we should compute them.
But for the first point, to me the MNI/realistic is the safer default:
- If you use idealized/spherical without realizing it, you'll (in theory at least?) get worse source localization and probably not know/notice, but your viz will look fine
- If you use MNI/realistic without realizing it, you'll get viz that you don't quite reconize and wonder why it looks funny, but your source loc will be better
To me the downside/problem in the second case is less bad/risky than the first.
As for the second point -- whether to compute on the fly -- let's not rehash here and continue discussion here instead:
https://github.com/mne-tools/mne-python/pull/8753#issuecomment-761045289
Even if we have a way to compute realistic positions, we can think of the problem the same way I think.
Currently, there is no way to tell in which category a given montage belongs to
See https://github.com/mne-tools/mne-python/issues/10747#issuecomment-1153967900 for a proposal/issue to help with this
I think the first step would be to clearly state in each montage's description what the montage is based on (idealized, or realistic coordinates), and whether it's a good or a maybe not so good idea to use it for source modeling.
As I quoted above, EEGLAB has also switched to MNI-based montages when doing dipole fits:
As of 2021, the default channel location file for electrode position is the MNI file, which is best suited for source localization. Before 2021, it was the BESA spherical location file.
Agreed, that would be a great first step. Bonus points for putting in the code somewhere what the original source was if possible, too. It might require some deep investigative work, though :)
@larsoner exactly, both use cases are important I think, so by documenting which montage is realistic and which is template we would already improve things a lot.
And yes, let's discuss adding on-the-fly calculations in #7472! It would only work for spherical models, yes. All other montages can remain in pre-computed (measured) files. Currently, we don't even have calculated 10-20 locations I think – I always use biosemi64 for this reason (which is computed from a sphere).
I always use biosemi64 for this reason (which is computed from a sphere).
Same! This one or easycap-M1
@larsoner I agree with the default montage tradeoffs you describe but I'm not sure if it is possible to perform source localization without noticing what kind of montage you have. You still have to coregister channel positions to the MRI (even if its the template), so you should be able to notice your channels are spherical, not realistic. Also, for their first source localization people would very likely follow some tutorial where we could highlight the benefit of realistic channel positions.
I'm not sure if it is possible to perform source localization without noticing what kind of montage you have
I think you ~~overestimate~~ underestimate how easy it is for people not to understand what an incorrect coreg looks like...
You still have to coregister channel positions to the MRI (even if its the template)
Typically people do not need to actually. fsaverage is already in MNI space, and we support trans='fsaverage'. In our existing tutorials this "just works" and I expect people would expect this to continue to be the case
I'm not sure if it is possible to perform source localization without noticing what kind of montage you have
I think you overestimate how easy it is for people not to understand what an incorrect coreg looks like...
I believe I've used a spherical montage together with trans='fsaverage' in the past myself, so … yes. :)
Ok, I am convinced. :)