PCA loadings matrix not quite orthogonal, benchmarks indicate errors.
Problem
PCA is giving me very different values for the loadings matrix when compared with CPU implementations of pca. I have compared the algorithm with fbpca and two of sklearn's implementations of PCA (randomized and full) which return identical values. Additionally I have found that transforming the data using the loadings matrix P gives different results when compared to T and that (at odds with T) the resulting columns are not quite (but almost) orthogonal.
Confusingly, when using PCA in a machine learning pipeline (and passing back the loadings matrix P to transform holdout data) I get scores almost identical to the aforementioned implementations. I expect this is due to P being not quite orthogonal. Oddly, the algorithm has clearly converged.
I am passing the data into PCA as per the example as follows:
Yp = GPUArray((m, n), Y.dtype, order='F')
Yp.set(Y)
pca = PCA(n_components=3)
T = pca.fit_transform(Yp)
In an effort to better understand the problem I have been playing with the svd implementation and have found some very odd behavior. The code and the output are self explanatory but suffice it to say that passing in column major matrices appears to be the root cause. However this is required as svd now requires a transposed matrix or will error out (fyi the example is broken because of this). Perhaps this is unrelated.
linalg.init()
Yg = GPUArray((n, m), Y.dtype, order='F')`
Yg.set(Y.T.copy())
Ug, Sg, Vg = gsvd(Yg, 'S', 'S')
print(Ug.shape)
print(Sg.shape)
print(Vg.shape)
Ug.T
(3, 3) (3,) (3, 5) array([[-0.74217098, -0.55579189, -0.37453653], [ 0.66994212, -0.63103686, -0.39111384], [-0.01896846, -0.54119114, 0.84068564]])
U, S, V = csvd(Y, full_matrices=False)
print(U.shape)
print(S.shape)
print(V.shape)
V
(5, 3) (3,) (3, 3) array([[-0.74217098, -0.55579189, -0.37453653], [-0.66994212, 0.63103686, 0.39111384], [-0.01896846, -0.54119114, 0.84068564]])
I'm scratching my head over this having experimented quite a bit and reviewing the referenced literature (I can find no bugs in the implementation). My only conclusion is that something is awry in how I pass in my data to be column major. However since (unlike the svd) the shapes of T, P, and U seem to be in order I am at a loss for ideas. Any insight is greatly appreciated as the algorithm performs extremely well in terms of speed and will be very useful! Thanks for putting this together :).
Environment
- Ubuntu 18.04
- Python 3.6.6 (Anaconda)
- Anaconda Cuda
- PyCUDA 2018.1.1
- scikit-cuda master (cloned)
Chris, I cannot follow you. Why do you expect T, the principal components, to have orthonormal columns? PCA imposes only constraints on the weight vectors (I guess that is what you are calling loadings), which turn out to be the eigenvectors of the covariance matrix.
Can you print the values for T?
Best, Ben
Does the PCA demo work for you? It still works fine for me. You didn't show what happened with the PCA sample you have there so I am not sure what's wrong with your output. The F-ordered arrays is required for the cublas functions. That's the only reason for that.
Hey guys, I believe I have isolated the problem. It appears that the differences I am seeing are a limitation of the algorithm and not a bug. To gain intuition I ran PCA using four different implementations: The GPU algorithm in question, sklearn's PCA using the full svd and a randomized algorithm, and facebook's implementation of PCA.
I found that in general when an algorithm could return the transformed matrix directly the dot product of the first and second components were essentially zero as in the example (fb PCA was the only implementation for which the transformed matrix was not returned directly). However when performing the same task but transforming the data using the loadings matrix each algorithm had a varying degree of success maintaining orthogonality. In general (without a careful statistical comparison) I found orthogonality was maintained best by the algorithms in the following order, where best is first:
- sklearn full svd
- sklearn randomized
- Fb
- GPU
For the GPU implementation I tried to push the tolerance down and max iterations up but this didn't have much effect, nor would it be particularly advantageous computationally. Hence the "nearly correct" scores I found when benchmarking the GPU algorithm in a machine learning pipeline make perfect sense. An example run of the code (motivated by the example) is as follows:
import pycuda.autoinit
import pycuda.gpuarray as gpuarray
import numpy as np
import skcuda.linalg as linalg
from skcuda.linalg import PCA as cuPCA
X = np.random.rand(1000, 100)
X = X - X.mean(axis=0)
m, n = X.shape
pca = cuPCA(n_components=4, epsilon=1e-50, max_iter=10**5)
X_gpu = gpuarray.GPUArray(X.shape, np.float64, order="F")
X_gpu.set(X)
T_gpu, P_gpu = pca.fit_transform(X_gpu)
T_gpu_derived = X @ P_gpu.get()
linalg.dot(T_gpu[:,0], T_gpu[:,1])
-1.1102230246251565e-15
from sklearn.decomposition import PCA
pca = PCA(n_components=4, svd_solver='full')
T_cpu = pca.fit_transform(X)
T_cpu_derived = X @ pca.components_.T
np.dot(T_cpu[:,0], T_cpu[:,1])
-3.352873534367973e-14
pca_randomized = PCA(n_components=4, svd_solver='randomized')
T_randomized = pca_randomized.fit_transform(X)
T_randomized_derived = X @ pca_randomized.components_.T
np.dot(T_randomized[:,0], T_randomized[:,1])
1.3322676295501878e-15
from fbpca import pca
U_fb, s_fb, Va_fb = pca(X, k=4, raw=True)
T_fb = X @ Va_fb.T
print('Gpu ' + str(np.dot(T_gpu_derived[:,0], T_gpu_derived[:,1])))
print('Cpu full ' + str(np.dot(T_cpu_derived[:,0], T_cpu_derived[:,1])))
print('Cpu randomized ' + str(np.dot(T_randomized_derived[:,0], T_randomized_derived[:,1])))
print('Fb ' + str(np.dot(T_fb[:,0], T_fb[:,1])))
Gpu 1.383103781345087 Cpu full 3.863576125695545e-14 Cpu randomized -0.03190332182138622 Fb 0.21848553288253514
Thanks for the help!
I have tried the example of the documentation and the results are orthogonal. The dot product is: 9.094947017729282e-12
However, the projections that PCA makes doesn't look fine. Below is the MWE of applying PCA to the iris dataset.
import pycuda.autoinit
import pycuda.gpuarray as gpuarray
import numpy as np
import skcuda.linalg as linalg
from skcuda.linalg import PCA as cuPCA
import matplotlib.pyplot as plt
from sklearn import datasets
iris = datasets.load_iris()
X = iris.data
y = iris.target
pca = cuPCA(n_components=2)
X_gpu = gpuarray.GPUArray((150,4), np.float64, order="F")
X_gpu.set(X)
T_gpu = pca.fit_transform(X_gpu)
print(linalg.dot(T_gpu[:,0], T_gpu[:,1]))
T = np.array(T_gpu.get())
plt.scatter(T[:,0], T[:,1], c=y, cmap=plt.cm.Set1, edgecolor='k', s=20)
plt.show()
The result is the following, and it is not similar with the projections of PCA that offers sklearn, Weka, R or Matlab.
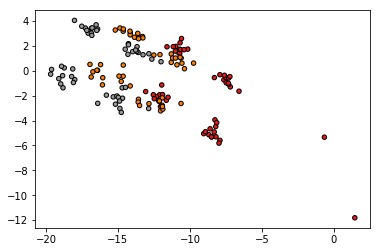
Is there any problem with my code or with the PCA implementation?
I have fixed this issue. If you look at the relevant pull request #283 you can see the combination of errors that made the code pass the dot product tests, but fail at producing meaningful projections. Thank you all for pointing out these hard-to-track issues!