gmx_MMPBSA: step3,awk: cmd. line:468: (FILENAME=_pid.pdb FNR=119376) warning: exp: argument 886.554 is out of range
Dear Dr. Li, I have used gmx_MMPBSA.bsh to calculate the Gibes free energy. But the last step always reported error which is :awk: cmd. line:468: (FILENAME=_pid.pdb FNR=119376) warning: exp: argument 886.554 is out of range. I have checked the .tpr, .xtc, index.ndx file , and did not find any problems. I don't know how to solve this problem, please help me.
For exponential function exp(x), if x is out of range (valid range is about |x|<700), exp(x) of awk will return an error. This is what you got. This error can be fixed easily, but it indicates there are some issues or errors of the calculation, so I will not fix it before I am sure the calculation is correct. Please check the values you got, if the results are reliable, I will fix this issue.
For exponential function
exp(x), if x is out of range (valid range is about |x|<700),exp(x)of awk will return an error. This is what you got. This error can be fixed easily, but it indicates there are some issues or errors of the calculation, so I will not fix it before I am sure the calculation is correct. Please check the values you got, if the results are reliable, I will fix this issue.
I am not sure whether the result is reliable, so I copy them to here. This is the content of _pid~MMPBSA.dat. Besides, I found that the CAS.pdb file not generate successfully. I wonder if this will affect the result. #Frame Binding( with DH ) | MM ( with DH ) PB SA | COU ( with DH ) VDW | PBcom PBpro PBlig | SAcom SApro SAlig _pid~0ns -87789.926(-12325.654) | -87789.926(-12325.654) 0.000 0.000 | -85266.073(-9801.801) -2523.853 | 0.000 0.000 0.000 | 0.000 0.000 0.000 _pid~1ns -87280.285(-12914.850) | -87280.285(-12914.850) 0.000 0.000 | -84401.225(-10035.790) -2879.061 | 0.000 0.000 0.000 | 0.000 0.000 0.000 _pid~2ns -90178.816(-13565.068) | -90178.816(-13565.068) 0.000 0.000 | -87171.834(-10558.085) -3006.982 | 0.000 0.000 0.000 | 0.000 0.000 0.000 _pid~3ns -91439.790(-13960.454) | -91439.790(-13960.454) 0.000 0.000 | -88300.599(-10821.262) -3139.191 | 0.000 0.000 0.000 | 0.000 0.000 0.000 _pid~4ns -90794.753(-13902.319) | -90794.753(-13902.319) 0.000 0.000 | -87617.569(-10725.134) -3177.184 | 0.000 0.000 0.000 | 0.000 0.000 0.000 _pid~5ns -92442.358(-14821.547) | -92442.358(-14821.547) 0.000 0.000 | -89080.942(-11460.131) -3361.416 | 0.000 0.000 0.000 | 0.000 0.000 0.000 ----------------------------------|------------------------------------------|--------------------------------| dH= -89987.655(-13581.648) | -89987.655(-13581.648) 0.000 0.000 | -86973.040(-10567.034) -3014.615 |
| -TdS= +inf( 1251.553) |
|---|
| dG= +inf(-12330.095) kJ/mol = +inf(-2946.964) kcal/mol |
| Ki= +INF(0.000E+00) uM = +INF(0.000E+00) nM |
| +==============================================================+ |
dG = RTln(Ki) = -RTln(KA)
Ki = 1/KA = exp(dG/RT) = IC50 = EC50
+==============================================================+ | M(mol/L) | m/u/pM | dG(kcal/mol) | dG(kJ/mol) | Affinity | |==============================================================| | 0.1 | 100 mM | -1.364 | -5.708 | | | 0.01 | 10 mM | -2.728 | -11.416 | | | 0.001 | 1 mM | -4.093 | -17.124 | | | 0.0001 | 100 uM | -5.457 | -22.832 | Weak | |--------------------------------------------------------------| | 0.00001 | 10 uM | -6.821 | -28.540 | | | 1.0E-06 | 1 uM | -8.185 | -34.248 | Medium | |--------------------------------------------------------------| | 1.0E-07 | 100 nM | -9.550 | -39.956 | | | 1.0E-08 | 10 nM | -10.914 | -45.664 | | | 1.0E-09 | 1 nM | -12.278 | -51.372 | Strong | |--------------------------------------------------------------| | 1.0E-10 | 100 pM | -13.642 | -57.080 | | | 1.0E-11 | 10 pM | -15.007 | -62.788 | | | 1.0E-12 | 1 pM | -16.371 | -68.496 | Very strong | +==============================================================+
#Frame Binding( with DH ) | MM ( with DH ) PB SA | COU ( with DH ) VDW | PBcom PBpro PBlig | SAcom SApro SAlig _pid~0ns -87789.926(-12325.654) | -87789.926(-12325.654) 0.000 0.000 | -85266.073(-9801.801) -2523.853 | 0.000 0.000 0.000 | 0.000 0.000 0.000 _pid~1ns -87280.285(-12914.850) | -87280.285(-12914.850) 0.000 0.000 | -84401.225(-10035.790) -2879.061 | 0.000 0.000 0.000 | 0.000 0.000 0.000 _pid~2ns -90178.816(-13565.068) | -90178.816(-13565.068) 0.000 0.000 | -87171.834(-10558.085) -3006.982 | 0.000 0.000 0.000 | 0.000 0.000 0.000 _pid~3ns -91439.790(-13960.454) | -91439.790(-13960.454) 0.000 0.000 | -88300.599(-10821.262) -3139.191 | 0.000 0.000 0.000 | 0.000 0.000 0.000 _pid~4ns -90794.753(-13902.319) | -90794.753(-13902.319) 0.000 0.000 | -87617.569(-10725.134) -3177.184 | 0.000 0.000 0.000 | 0.000 0.000 0.000 _pid~5ns -92442.358(-14821.547) | -92442.358(-14821.547) 0.000 0.000 | -89080.942(-11460.131) -3361.416 | 0.000 0.000 0.000 | 0.000 0.000 0.000 ----------------------------------|------------------------------------------|--------------------------------| dH= -89987.655(-13581.648) | -89987.655(-13581.648) 0.000 0.000 | -86973.040(-10567.034) -3014.615 |
| -TdS= +inf( 1251.553) |
|---|
| dG= +inf(-12330.095) kJ/mol = +inf(-2946.964) kcal/mol |
| Ki= +INF(0.000E+00) uM = +INF(0.000E+00) nM |
| +==============================================================+ |
dG = RTln(Ki) = -RTln(KA)
Ki = 1/KA = exp(dG/RT) = IC50 = EC50
+==============================================================+ | M(mol/L) | m/u/pM | dG(kcal/mol) | dG(kJ/mol) | Affinity | |==============================================================| | 0.1 | 100 mM | -1.364 | -5.708 | | | 0.01 | 10 mM | -2.728 | -11.416 | | | 0.001 | 1 mM | -4.093 | -17.124 | | | 0.0001 | 100 uM | -5.457 | -22.832 | Weak | |--------------------------------------------------------------| | 0.00001 | 10 uM | -6.821 | -28.540 | | | 1.0E-06 | 1 uM | -8.185 | -34.248 | Medium | |--------------------------------------------------------------| | 1.0E-07 | 100 nM | -9.550 | -39.956 | | | 1.0E-08 | 10 nM | -10.914 | -45.664 | | | 1.0E-09 | 1 nM | -12.278 | -51.372 | Strong | |--------------------------------------------------------------| | 1.0E-10 | 100 pM | -13.642 | -57.080 | | | 1.0E-11 | 10 pM | -15.007 | -62.788 | | | 1.0E-12 | 1 pM | -16.371 | -68.496 | Very strong | +==============================================================+
#Frame Binding( with DH ) | MM ( with DH ) PB SA | COU ( with DH ) VDW | PBcom PBpro PBlig | SAcom SApro SAlig _pid~0ns -87789.926(-12325.654) | -87789.926(-12325.654) 0.000 0.000 | -85266.073(-9801.801) -2523.853 | 0.000 0.000 0.000 | 0.000 0.000 0.000 _pid~1ns -87280.285(-12914.850) | -87280.285(-12914.850) 0.000 0.000 | -84401.225(-10035.790) -2879.061 | 0.000 0.000 0.000 | 0.000 0.000 0.000 _pid~2ns -90178.816(-13565.068) | -90178.816(-13565.068) 0.000 0.000 | -87171.834(-10558.085) -3006.982 | 0.000 0.000 0.000 | 0.000 0.000 0.000 _pid~3ns -91439.790(-13960.454) | -91439.790(-13960.454) 0.000 0.000 | -88300.599(-10821.262) -3139.191 | 0.000 0.000 0.000 | 0.000 0.000 0.000 _pid~4ns -90794.753(-13902.319) | -90794.753(-13902.319) 0.000 0.000 | -87617.569(-10725.134) -3177.184 | 0.000 0.000 0.000 | 0.000 0.000 0.000 _pid~5ns -92442.358(-14821.547) | -92442.358(-14821.547) 0.000 0.000 | -89080.942(-11460.131) -3361.416 | 0.000 0.000 0.000 | 0.000 0.000 0.000 ----------------------------------|------------------------------------------|--------------------------------| dH= -89987.655(-13581.648) | -89987.655(-13581.648) 0.000 0.000 | -86973.040(-10567.034) -3014.615 |
| -TdS= +inf( 1251.553) |
|---|
| dG= +inf(-12330.095) kJ/mol = +inf(-2946.964) kcal/mol |
| Ki= +INF(0.000E+00) uM = +INF(0.000E+00) nM |
| +==============================================================+ |
dG = RTln(Ki) = -RTln(KA)
Ki = 1/KA = exp(dG/RT) = IC50 = EC50
+==============================================================+ | M(mol/L) | m/u/pM | dG(kcal/mol) | dG(kJ/mol) | Affinity | |==============================================================| | 0.1 | 100 mM | -1.364 | -5.708 | | | 0.01 | 10 mM | -2.728 | -11.416 | | | 0.001 | 1 mM | -4.093 | -17.124 | | | 0.0001 | 100 uM | -5.457 | -22.832 | Weak | |--------------------------------------------------------------| | 0.00001 | 10 uM | -6.821 | -28.540 | | | 1.0E-06 | 1 uM | -8.185 | -34.248 | Medium | |--------------------------------------------------------------| | 1.0E-07 | 100 nM | -9.550 | -39.956 | | | 1.0E-08 | 10 nM | -10.914 | -45.664 | | | 1.0E-09 | 1 nM | -12.278 | -51.372 | Strong | |--------------------------------------------------------------| | 1.0E-10 | 100 pM | -13.642 | -57.080 | | | 1.0E-11 | 10 pM | -15.007 | -62.788 | | | 1.0E-12 | 1 pM | -16.371 | -68.496 | Very strong | +==============================================================+
There's no PB or SA value, so the calculaiton was not correct. Please reproduce the example before run on your own systems.
I have already run the example ,the result also has no PB or SA value, as below. #Frame Binding( with DH ) | MM ( with DH ) PB SA | COU ( with DH ) VDW | PBcom PBpro PBlig | SAcom SApro SAlig _pid~0ns -468.207( -462.925) | -468.207( -462.925) 0.000 0.000 | -147.073( -141.791) -321.134 | 0.000 0.000 0.000 | 0.000 0.000 0.000 _pid~1ns -488.042( -478.346) | -488.042( -478.346) 0.000 0.000 | -176.306( -166.609) -311.736 | 0.000 0.000 0.000 | 0.000 0.000 0.000 _pid~2ns -484.874( -481.025) | -484.874( -481.025) 0.000 0.000 | -146.223( -142.374) -338.651 | 0.000 0.000 0.000 | 0.000 0.000 0.000 _pid~3ns -555.573( -550.681) | -555.573( -550.681) 0.000 0.000 | -179.018( -174.126) -376.555 | 0.000 0.000 0.000 | 0.000 0.000 0.000 _pid~4ns -474.792( -471.219) | -474.792( -471.219) 0.000 0.000 | -144.988( -141.415) -329.804 | 0.000 0.000 0.000 | 0.000 0.000 0.000 _pid~5ns -480.970( -476.171) | -480.970( -476.171) 0.000 0.000 | -162.986( -158.187) -317.984 | 0.000 0.000 0.000 | 0.000 0.000 0.000 _pid~6ns -513.379( -500.065) | -513.379( -500.065) 0.000 0.000 | -181.466( -168.152) -331.913 | 0.000 0.000 0.000 | 0.000 0.000 0.000 _pid~7ns -505.047( -493.464) | -505.047( -493.464) 0.000 0.000 | -170.461( -158.878) -334.585 | 0.000 0.000 0.000 | 0.000 0.000 #Frame Binding( with DH ) | MM ( with DH ) PB SA | COU ( with DH ) VDW | PBcom PBpro PBlig | SAcom SApro SAlig _pid~0ns -468.207( -462.925) | -468.207( -462.925) 0.000 0.000 | -147.073( -141.791) -321.134 | 0.000 0.000 0.000 | 0.000 0.000 0.000 _pid~1ns -488.042( -478.346) | -488.042( -478.346) 0.000 0.000 | -176.306( -166.609) -311.736 | 0.000 0.000 0.000 | 0.000 0.000 0.000 _pid~2ns -484.874( -481.025) | -484.874( -481.025) 0.000 0.000 | -146.223( -142.374) -338.651 | 0.000 0.000 0.000 | 0.000 0.000 0.000 _pid~3ns -555.573( -550.681) | -555.573( -550.681) 0.000 0.000 | -179.018( -174.126) -376.555 | 0.000 0.000 0.000 | 0.000 0.000 0.000 _pid~4ns -474.792( -471.219) | -474.792( -471.219) 0.000 0.000 | -144.988( -141.415) -329.804 | 0.000 0.000 0.000 | 0.000 0.000 0.000 _pid~5ns -480.970( -476.171) | -480.970( -476.171) 0.000 0.000 | -162.986( -158.187) -317.984 | 0.000 0.000 0.000 | 0.000 0.000 0.000 _pid~6ns -513.379( -500.065) | -513.379( -500.065) 0.000 0.000 | -181.466( -168.152) -331.913 | 0.000 0.000 0.000 | 0.000 0.000 0.000 _pid~7ns -505.047( -493.464) | -505.047( -493.464) 0.000 0.000 | -170.461( -158.878) -334.585 | 0.000 0.000 0.000 | 0.000 0.000 0.000 _pid~8ns -453.624( -449.163) | -453.624( -449.163) 0.000 0.000 | -134.994( -130.533) -318.630 | 0.000 0.000 0.000 | 0.000 0.000 0.000 _pid~9ns -513.094( -507.992) | -513.094( -507.992) 0.000 0.000 | -156.809( -151.707) -356.285 | 0.000 0.000 0.000 | 0.000 0.000 0.000 _pid~10ns -467.055( -463.589) | -467.055( -463.589) 0.000 0.000 | -140.663( -137.196) -326.393 | 0.000 0.000 0.000 | 0.000 0.000 0.000 ----------------------------------|------------------------------------------|--------------------------------| dH= -491.332( -484.967) | -491.332( -484.967) 0.000 0.000 | -158.272( -151.906) -333.061 |
| -TdS= 31.782( 29.877) |
|---|
| dG= -459.550( -455.090) kJ/mol = -109.835( -108.769) kcal/mol |
| Ki= 3.093E-75(1.869E-74) uM = 3.093E-72(1.869E-71) nM |
| +==============================================================+ |
dG = RTln(Ki) = -RTln(KA)
Ki = 1/KA = exp(dG/RT) = IC50 = EC50
+==============================================================+ | M(mol/L) | m/u/pM | dG(kcal/mol) | dG(kJ/mol) | Affinity | |==============================================================| | 0.1 | 100 mM | -1.364 | -5.708 | | | 0.01 | 10 mM | -2.728 | -11.416 | | | 0.001 | 1 mM | -4.093 | -17.124 | | | 0.0001 | 100 uM | -5.457 | -22.832 | Weak | |--------------------------------------------------------------| | 0.00001 | 10 uM | -6.821 | -28.540 | | | 1.0E-06 | 1 uM | -8.185 | -34.248 | Medium | |--------------------------------------------------------------| | 1.0E-07 | 100 nM | -9.550 | -39.956 | | | 1.0E-08 | 10 nM | -10.914 | -45.664 | | | 1.0E-09 | 1 nM | -12.278 | -51.372 | Strong | |--------------------------------------------------------------| | 1.0E-10 | 100 pM | -13.642 | -57.080 | | | 1.0E-11 | 10 pM | -15.007 | -62.788 | | | 1.0E-12 | 1 pM | -16.371 | -68.496 | Very strong | +==============================================================+
There's no PB or SA value, so the calculaiton was not correct. Please reproduce the example before run on your own systems.
Dear Dr. Li, I have reproduced the example of gmx_MMPBSA on my computer, there is no error report in log file. But the values of PB, SA are also zero. I wonder which step was wrong. Please tell me the right way to calculate ΔG. Thank you very much!
I told you there are no values for PB and SA, that's the problem. The PB, SA values are calculated by APBS, it means you have to install APBS on your machine first and then set the APBS variable correctly in the script. If you do not understand what I am talking about, read the documents first.
Dear Dr. Li,
I have reperform the free energy calculation by resetting the APBS. Now I can get the value of PB and SA, but I can not get the value of total free energy change! I wonder the reason, so I need your help. Here is the output and the error report .
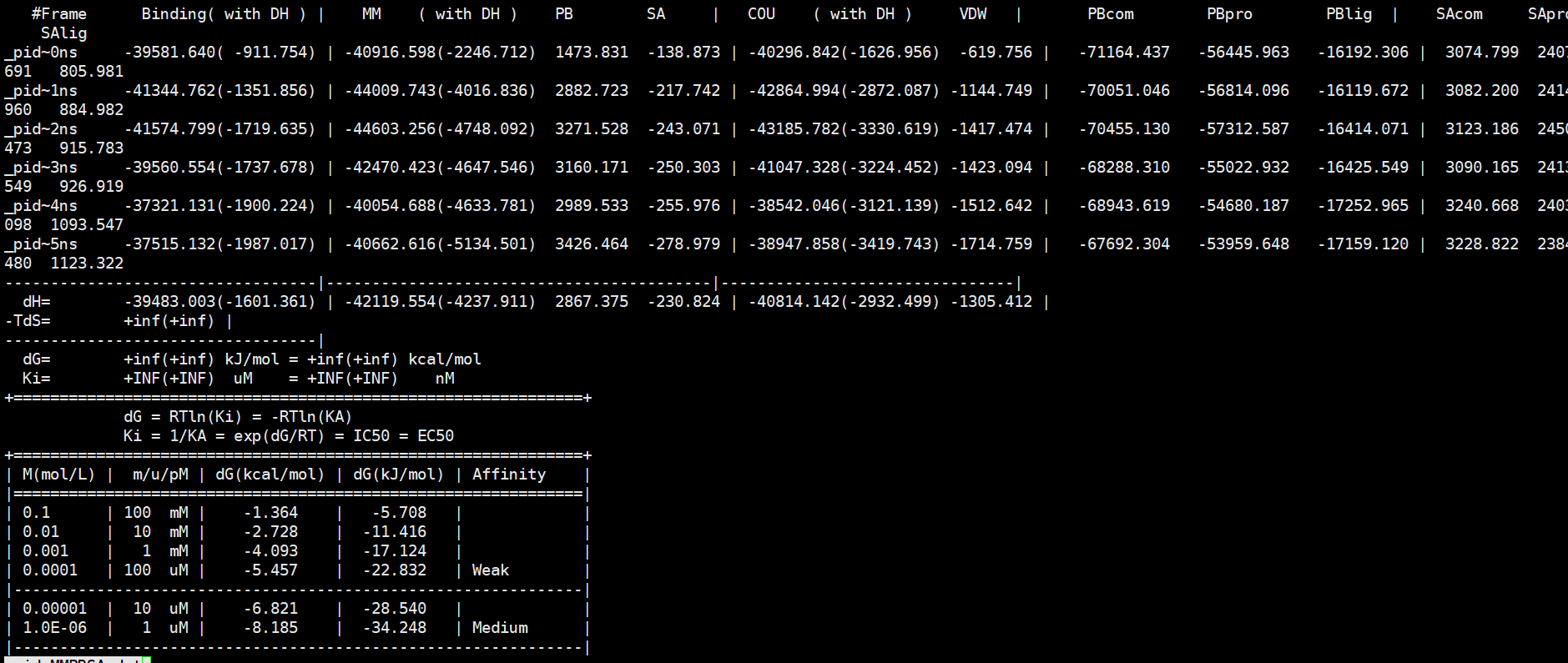

I did not see obvious errors for the numbers, but the COU and VDW values are quite large, either there are bad structures or the net charges are large. Please make sure the structures you used to run MMPBSA calculations are equilibrated well. If there are large net charges in the system, e.g. DNA, we can do nothing. Remember, there are some limitations for MMPBSA method.
I can change the script to show non-inf, but that does not solve the problem.
Dear Dr. Li, In our system, there is a DNA exist, so as you say, the value is infinite exactly because of this! That is so sad! I wonder if there is other ways to calculate such system with DNA?
It's still an open question that how to perform MMPBSA calculations for highly charged systems, you can check some papers for this topic. I have some idea about this topic, but it needs validations, so I can't put them in the script simply.
For your system, maybe you can try:
- increase the dielectric epsilon from 2 to 4 or even large values, this will decrease the COU and PB values
- for DNA, if it's highly charged, it's not a good idea to use it alone in MMPBSA calculations,. Try to include the ions binding to DNA as part of the system.
- maybe it is also a good idea to include some water as part of the system.