GTDBTk
 GTDBTk copied to clipboard
GTDBTk copied to clipboard
GTDB-Tk: a toolkit for assigning objective taxonomic classifications to bacterial and archaeal genomes.
## Environment - [x] Using a conda environment (include the output of `conda list && conda list --revisions`) conda list ``` # packages in environment at /mnt/mibi_ngs/keerthi/miniconda3/envs/gtdbtk-2.1.0: # # Name...
## Environment - [ ] Installed via pip (include the output of `pip list`) - [X] Using a conda environment (include the output of `conda list && conda list --revisions`)...
Dear, Can you help me? Please see below: The first error was that several Tigrfam and pfam files (markers) were missing from the gtdbk r207 database. So I include them,...
Hi @donovan-parks! I just ran into an error while running the gtdbtk classify_wf command. My command was: **gtdbtk classify_wf --genome_dir /lustre/rsharma/FLOOR-METAGENOME-NOVASEQ/ASSEMBLY/MAXBIN_REZ/GOOD_QUALITY_BINS/GBINS/ --out_dir /lustre/rsharma/FLOOR-METAGENOME-NOVASEQ/GTDBTK_REZ/** Terminal output: 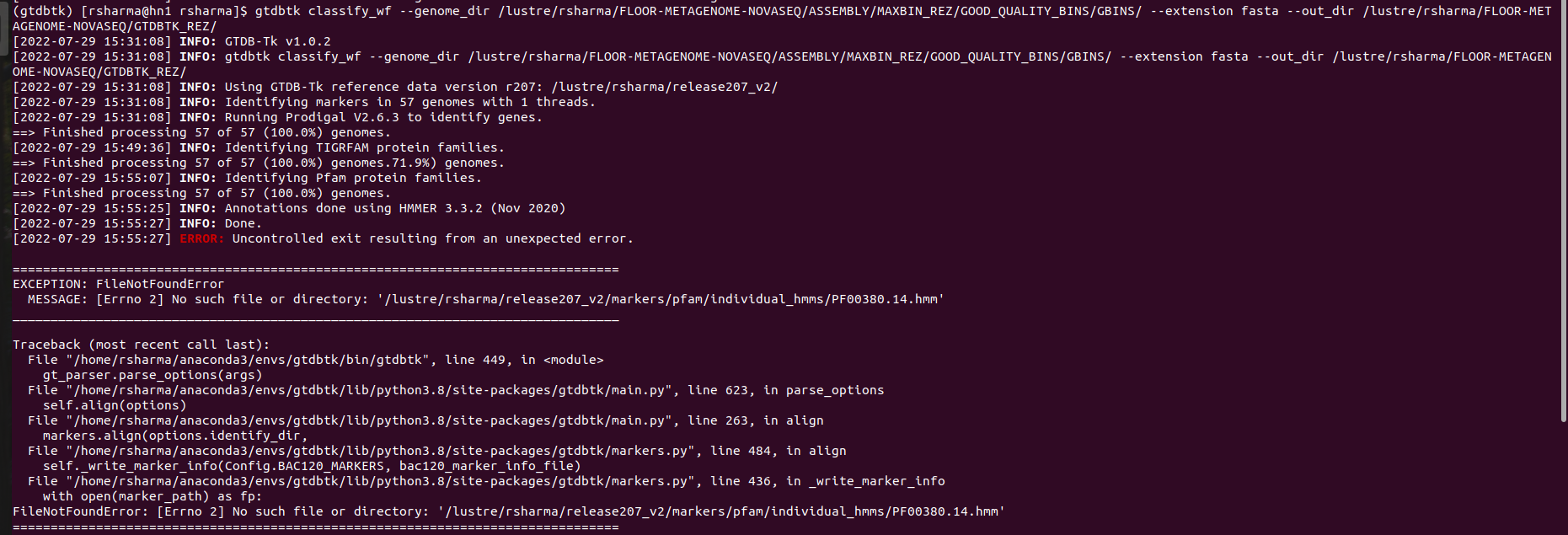 ERROR:...
Add a line in the log file listing how many genomes have a warning. Similar to `[2022-07-12 12:00:00] WARNING: 10 out of 100 genomes ( 10% ) have a warning,...
I would like to ask that how we suppose to know how much reads or percentage of reads were classified? I would like to compare it to other analysis method.
I had some bad experiences with gtdb-tk classify which I run on large datasets. - For example, It passed all the steps but then failed because the symlinked files already...
It would be good if users could supply genomes as called genes in amino acid space. Basically just a flag (--genes) to indicate the FASTA file contains genes and then...
Thanks for the great tool! This suggestion is definitely not critical, but something I think could be useful. I definitely like that on the GTDB website you can search for...
Hi there I used GTDB-tk (v1.4) to generate MSA and then used it to construct a phylogenetic tree. However, IQ-tree reported that some of the sequences have >50% gaps/ambiguity. As...
