A case against is_binned
Hi everyone,
I apologize in advance that this is going to be a long post, but I would like to advocate for a somewhat significant change in hyperspy, which I think deserves thorough justification. I would also like to state that my statement should not be taken as anything else as genuine effort to improve hyperspy for everyone after deep study of the code base.
Over time, I have developed increasing skepticism towards the usefulness of the whole binned and unbinned axes paradigm. Specifically, I have the following issues with them:
0 - The concept of binned axis is an odd, hyperspy-specific thing.
- I Never seen this concept as an axis property in data analysis (software or literature).
- It is an added step in the learning curve which might discourage new users, and stay puzzling even to the most experienced.
1 - Binned signals are badly documented
- from doc: "[...] histogram of a probability distribution function should have
is_binnedset to True". Most datasets can fit in this definition yet few are binned. What would be counter examples? Also, the link with a probability distribution function can be misleading (see histogram example). - from doc: "[...] some methods operate differently on signals that are binned". We don't know which ones, why it is necessary, what does it change for the user.
- The documentation does not link to scientific literature on analysis of binned or unbinned data. Relevant information is difficult to find due to the many usages of "binning" in science to denote different things that are vaguely related but not identical.
2 - is_binned gives opaque behaviour to integrate1D. From doc: "For binned axes, the detector already provides the per-channel integration of the signal. Therefore, in this case, integrate1D performs a simple summation along the given axis. In contrast, for unbinned axes, integrate1D calls the integrate_simpson method."
- As I understand it, the idea is that
integrate1Dreturns an output dimension of let's say [Counts] regardless of whether the dimension is [Counts/eV] or [Counts] at the method's input (for an eV signal axis). This does not relate to the data being a probability distribution function, but how it is normalized. This is already problematic to the definition ofis_binned. - I think
integrate1Dshould return the (simpson) integral, andsumreturn the sum. Both functions are clearly defined, and at runtime there should never be an ambiguity about which method applies. In other words, we should expect users to know what they want to achieve.
3 - is_binned gives hard to understand behaviour to fit:
- models fitted on binned signals are no longer true to the data. For example, when fitting a gaussian function to binned data, the "A" parameter is no longer the area under the curve. This behaviour is not documented or explained to the user.
- Fitting a signal along a binned axis gives the same result as dividing the signal by the axis gradient, setting
is_binned=Falseand then fitting the signal (see examples). Such renormalisation operation is rather trivial and we should expect users to know if they need to do it, why, when, and how. - In my opinion, any fitting module should always fit models true to the data or users will think the module is broken.
4 - Aside from the modifications to Integrate1D and fit, is_binned provide no other useful feature to hyperspy:
- I think this is explicit enough.
5 - The implementation and performance costs are large
- evaluating is_binned takes up to 40% of the execution time in multifit. At this point we're not sure it is going to be solved in v2.0 (see #3126 , #2949)
- In fact, the implementation of binned signals is not entirely done. e.g.
Model2D.__call__does not take into accountis_binned, yetbaseModel._linear_fithas an implementation that will impactModel2D.fit, possibly breaking it (haven't tried). #2427 - The implementation of any new model component must take
is_binnedinto account. - The implementation of new axes and signals must take
is_binnedinto account - maintaining is_binned requires a significant effort, for example in the upcoming v2.0 release, and will continue to require so.
- There remains bugs in is_binned (see example) and I am not sure its behaviour is correct for non uniform axes.
- It is only used in the context of EELSSpectra and EDSSpectra which are soon to be split from main hyperspy.
Now, how do I think we could make hyperspy better by removing is_binned:
0 - This would bring hyperspy closer to other data analysis software and python packages. Sense of familiarity is important to users and hyperspectral data analysis is difficult enough as it is, without introducing new data paradigms.
1 - The need to enhance the documentation on binned signal and educate users about them would no longer be needed. The need to better define why should an axis be binned or not would also no longer be needed. In other term, clarity is improved for users.
2 - There would be no ambiguity in code instances making use of Integrate1D, namely: EDSModel.get_line_intensity, EELSSpectrum.estimate_elastic_scattering_intensity, EELSSpectrum.estimate_thickness, EELSSpectrum.kramers_kronig_analysis, misc.eels.eels_constant. EELSSpectrum and EDSSpectrum are forced to is_binned=True at instantiation so integrate1D could be replaced by sum in these functions and that's it. In other words, there would not be impact of removing this aspect of Integrate1D and the inconsistency in its behaviour would be removed.
3 - Functions would always fit true to the data: evaluating fitted function on the data axis returns the best fit to data and parameters keep their meaning. If renormalization by the axis gradient is needed before fit, a warning could be raised to users so that they can consider creating a renormalized copy of the signal. Only EELSSpectra (possibly EDSSpectrum) would be affected and this could be handled at model creation time. Alternatively, behaviour of renormalizing axis could be integrated in the EELS model components which would also make the split easier. Implementation and maintenance of fit components would also be simpler.
4 - I see a large potential for simplification of the code base. fits would basically run twice as fast, new signals and components would be easier to implement.
to conclude, I think the is_binned paradigm has been developed to bring a solution to extremely specific analyses of EELS/EDS Signals (which I have no expertise in) and creates all sorts of inconveniences in other use cases. Now that hyperspy has an expanding user base and a variety of other applications than electron microscopy it has become a liability. What do you think? Would you be opened to such change?
Sorry again for being so straightforward in phrasing this, but I think this post is long enough and I really wanted to stay to the point. Hereafter, I will post a few examples to illustrate my point.
The first example concerns histograms. A signal with normal-distributed random data is created and its histogram fitted with a gaussian. The point of this example is to illustrate the behaviour of fitting and integrating binned signals in a case where the link with a probability distribution function is as clear as possible.
Example code:
import hyperspy.api as hs
from hyperspy._components.gaussian import Gaussian
import numpy as np
import matplotlib.pyplot as plt
s = hs.signals.Signal1D(np.random.randn(6666))
s = s.get_histogram(density=True)
# s.axes_manager.signal_axes[0].is_binned = False
print(f"histogram is_binned = {s.axes_manager.signal_axes[0].is_binned} \n")
m = s.create_model()
g = Gaussian(A=1000, centre=1240/400.0)
m.append(g)
m.fit()
m.print_current_values()
msig = m.as_signal()
res = msig.integrate1D(-1).data
print(f"integral1D result: {res}")
res2 = msig.sum(-1).data
print(f"sum result: {res2}")
res3 = msig.integrate_simpson(-1).data
print(f"integrate_simpson result: {res3}")
prompt result
histogram is_binned = True
Model1D: histogram
current_component_values: Gaussian
Active: True
Parameter Name | Free | Value | Std | Min | Max | Linear
============== | ======= | ========== | ========== | ========== | ========== | ======
A | True | 7.16679948 | 0.08175887 | 0.0 | None | True
centre | True | -0.0534630 | 0.01282962 | None | None | False
sigma | True | 0.97398851 | 0.01283114 | 0.0 | None | False
integral1D result: [7.16333975]
sum result: [7.16333975]
integrate_simpson result: [0.99389729]
prompt result when un-commenting the line setting is_binned = False
histogram is_binned = False
Model1D: histogram
current_component_values: Gaussian
Active: True
Parameter Name | Free | Value | Std | Min | Max | Linear
============== | ======= | ========== | ========== | ========== | ========== | ======
A | True | 0.99455927 | 0.01106550 | 0.0 | None | True
centre | True | -0.0985378 | 0.01257134 | None | None | False
sigma | True | 0.97852707 | 0.01257135 | 0.0 | None | False
integral1D result: [0.99453554]
sum result: [7.1249329]
integrate_simpson result: [0.99453554]
Here, the point is that it's very counterintuitive to fit a parameter A with a value different from 1 in a case where by construction it should be 1. The same could be said from the integral1D of the signal. Despite being the exact definition of a binned signal according to the docs, in this case clearly is_binned = False provides the correct behaviour
Here is a picture of the fitted signal
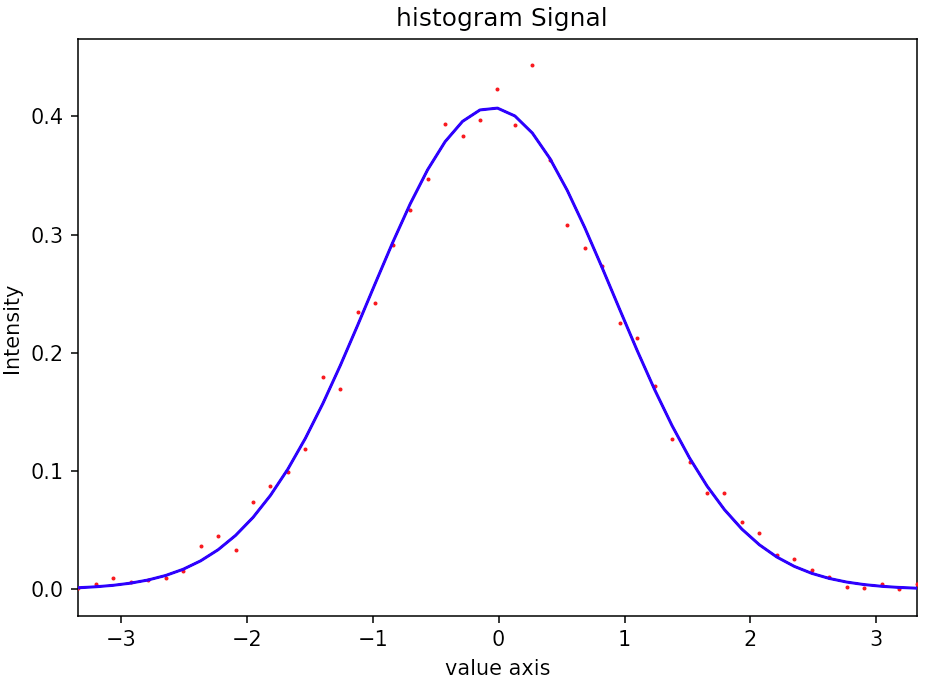
This second example shows the fitting of a core loss eels spectrum. The point of this example is to show the equivalence between fitting a renormalized signal with is_binned=False and a raw signal with is_binned=True in a non-trivial case.
from hyperspy.datasets import eelsdb
import numpy as np
s = eelsdb(title="Hexagonal Boron Nitride", spectrum_type="coreloss")[0]
s.add_elements(('B', 'N'))
s.set_microscope_parameters(beam_energy=300, convergence_angle=0.2, collection_angle=2.55)
ll = eelsdb(title="Hexagonal Boron Nitride", spectrum_type="lowloss")[0]
#First: intensities are kept as is and is_binned is True
m = s.create_model(ll=ll)
m.enable_fine_structure()
m.smart_fit()
print(f"Is_binned = {s.axes_manager[0].is_binned}.")
m.quantify()
#Second: intensities are renormalized and is_binned is False
s = s/np.gradient(s.axes_manager[0].axis)
s.axes_manager[0].is_binned = False
m = s.create_model(ll=ll)
m.enable_fine_structure()
m.smart_fit()
print(f"Is_binned = {s.axes_manager[0].is_binned}. Renormalized intensities")
m.quantify()
Which prompts the following:
Is_binned = True.
Absolute quantification:
Elem. Intensity
B 0.045648
N 0.048050
Is_binned = False. Renormalized intensities
Absolute quantification:
Elem. Intensity
B 0.045648
N 0.048050
My opinion is that in this case, the better option would be to additionally overload the eels model creator to detect if the axis is already in [eV^-1] units at creation and do the conversion if not while raising an informative warning. It can be verified easily that low-loss having is_binned=True or is_binned=False has no influence on the outcome
This final example shows that the behaviour of example 2 is reproduced on any other signal by fitting a Signal1D data type with a gaussian peak.
from hyperspy.datasets.artificial_data import get_luminescence_signal
from hyperspy._components.gaussian import Gaussian
import matplotlib.pyplot as plt
import numpy as np
#with is binned = True
s = get_luminescence_signal(uniform=True)
s.axes_manager[0].is_binned = True
m = s.create_model()
g = Gaussian(A=10, centre=400.0)
m.append(g)
m.fit()
m.print_current_values()
#Changing to is binned = False and renormalizing
s = s/np.gradient(s.axes_manager.signal_axes[0].axis)
s.axes_manager[0].is_binned = False
m = s.create_model()
m.append(g)
m.fit()
m.print_current_values()
m.plot()
plt.show()
Where the prompt is
Model1D: 0d-map Artificial Luminescence Signal
current_component_values: Gaussian
Active: True
Parameter Name | Free | Value | Std | Min | Max | Linear
============== | ======= | ========== | ========== | ========== | ========== | ======
A | True | 9269.35013 | 6.35386279 | 0.0 | None | True
centre | True | 375.002862 | 0.01979540 | None | None | False
sigma | True | 25.0095862 | 0.01979540 | 0.0 | None | False
Model1D: 0d-map Artificial Luminescence Signal
current_component_values: Gaussian
Active: True
Parameter Name | Free | Value | Std | Min | Max | Linear
============== | ======= | ========== | ========== | ========== | ========== | ======
A | True | 9269.35013 | 6.35386285 | 0.0 | None | True
centre | True | 375.002862 | 0.01979540 | None | None | False
sigma | True | 25.0095862 | 0.01979540 | 0.0 | None | False
Again, I would argue that in this case the user should know if the data needs renormalization before modelling.
Final example is more of a bug in the behaviour of eels and eds data types
from hyperspy.datasets.artificial_data import get_low_loss_eels_signal
s = get_low_loss_eels_signal()
s.axes_manager[0].is_binned = False
print(f"Axis has now is_binned attribute set to {s.axes_manager[0].is_binned}.")
s = s + 2
print(f"Axis has now is_binned attribute set to {s.axes_manager[0].is_binned}.")
The above code prompts
Axis has now is_binned attribute set to False.
Axis has now is_binned attribute set to True.
Because is_binned = True is hard-coded in the eels and eds signals creators, pretty much any operation on the data re-sets it to True.
This example illustrates both that it is unnecessary to cover both binned and unbinned axes in eels / eds signals (from the fact that it went unnoticed this far for what I know ^^), and that indeed it can be difficult to maintain is_binned property correctly :)
Now, I might absolutely be overlooking some important aspects of this feature.
Don't hesitate to signal me anything I get wrong on this. That being said, if you would agree to consider removing is_binned from future hyperspy version and implementing the changes I propose I think the split of eels and eds signals from the V2 would be the right opportunity.
Cheers
Nicolas
@Attolight-NTappy Thank you for the write up! This is maybe another case for the return of the HSEP for longer form issue and feature requests.
I'll wait for someone who has more experience with 1D Signals and signal fitting to maybe provide a defense. I'll say that I've always been confused by is_binned as well.
Mainly something like:
import hyperspy.api as hs
import numpy as np
s = hs.signals.Signal1D(np.ones((20,20,30)))
print(s.rebin(scale=(1,1,2)).axes_manager[2].is_binned) # prints False
Seems like it should print True but maybe I am misunderstanding what exactly we are trying to handle with the is_binned parameter.
All of that being said I am a huge supporter of tracking what modifications are done to some signal. I would almost prefer than all functions and their kwargs were tracked/ saved.
Pitching in as an extension maintainer (kikuchipy for electron backscatter diffraction): We do not use the attribute in question, so removing it would not break anything for us.