Extremely long hanging after `write_fst` for large files
Firstly, thanks for writing fst! I discovered this awesome package only recently and immediately like its speed!
ISSUE
As I start to replace the old saveRDS with write_fst, I discovered an issue: after running write_fst, if the file is very big, R hangs for many minutes (sometimes as long as 30mins) before being able to continue the next line. (Note: For smaller files, say 2-3GB after compression, there is no issue)
I regularly work with objects/files bigger than 50GB with >10,000 columns so I'm constantly saving them to disk. This is why I hit into the issue I mentioned.
As an aside: running read_fst() does not give this issue. Let's say there is a huge file (e.g. 50GB) I've already saved into fst format. After running read_fst(), R is able to run the next command immediately. BUT if I were to do write_fst, R hangs for some time.
SYSTEM SPEC
I'm not sure if the specs is the root cause, but here's the specs for the server I ran my scripts on:
- Debian 10
- 40-core VM with 900GB of RAM on Google Cloud (I've tried
write_fstwith 1 core, issue is still there) - R ver 3.3.3 (for compatibility reasons, I've not upgraded it)
- fst ver 0.9.0 (when it loads, it says it detects OpenMP and runs at 40 threads)
Hi @winston-p,
thanks for reporting your issue! The write times for fst files are expected to scale linearly with the number of columns and the row sizes, so it's certainly strange that the write operation becomes very slow when you cross a certain file size.
Could this be due to the way Google Cloud handles larger files? (I'm guessing memory won't be the issue on your system :-))
I don't have experience with VM's on Google Cloud, but could it be that larger files are split over multiple storage blobs?
You could check the existence of such a boundary by writing increasingly large files and see what happens, for example:
library(data.table)
library(ggplot2)
# function for generating sample data
sample_table <- function(nr_of_cols, nr_of_rows) {
as.data.table(
lapply(1:nr_of_cols, function(y) {
sample(1:1000, nr_of_rows, replace = TRUE)
})
)
}
# use increasing row numbers (you have to use bigger numbers here :-))
path <- tempdir()
row_sizes <- c(1e4, 1e5, 1e6, 1e7, 2.5e7)
# measure write times
res <- lapply(row_sizes, function(nr_of_rows) {
x <- sample_table(50, nr_of_rows)
y <- microbenchmark::microbenchmark(
fst::write_fst(x, paste0(path, "/", nr_of_rows, ".fst")),
times = 1
)
data.table(
SizeGB = 1e-9 * as.numeric(object.size(x)),
SpeedGBs = as.numeric(object.size(x)) / y$time)
})
On my system, this gives me:
# plot results
ggplot(rbindlist(res)) +
geom_line(aes(SizeGB, SpeedGBs)) +
geom_point(aes(SizeGB, SpeedGBs))
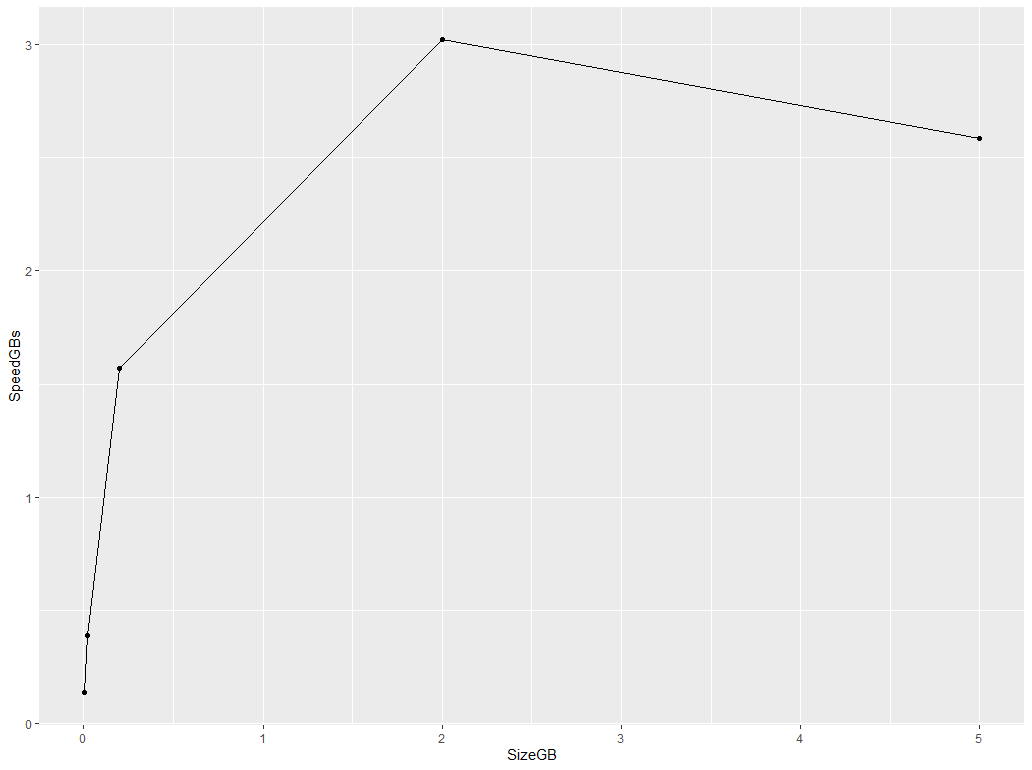
It would be interesting to see if you have a drop-off in speed after you cross a certain object size. Perhaps that will point us in the right direction as to why such a drop-off should exist...
thanks!
@MarcusKlik It's indeed rather weird that this is happening.
To clarify,, it's not that the write-speed becomes very slow with bigger file sizes. No, the write speed is still blazing fast. Writing into GCE's SSD, it writes at >0.5GB per sec before tapering off to 0.1GB per sec at the end, probably due to filling of write buffer (disclaimer: I'm not familiar with OS' lower-level workings).
The issue is with R hanging after write_fst has completed and R has moved to the next line. My R code is something like this:
write_fst(dt, "testfile.fst")
a <- 1L # Just to see when fst finishes writing
rm(dt); gc() # This is where R hangs for a really long time, sometimes up to 30min
I'll test on my own server later to see if this issue occurs due to Google Cloud or not. Will report back with my findings.
I noticed the same with windows before with large datasets . Do you have lots of strings? It might be R' gc at work.
On Thu., 2 Jan. 2020, 13:44 Winston P, [email protected] wrote:
@MarcusKlik https://github.com/MarcusKlik It's indeed rather weird that this is happening.
To clarify,, it's not that the write-speed becomes very slow with bigger file sizes. No, the write speed is still blazing fast. Writing into GCE's SSD, it writes at 0.5GB per sec before tapering off to 0.1GB per sec at the end, probably due to filling of write buffer (disclaimer: I'm not familiar with OS' lower-level workings).
The issue is with R hanging after write_fst has completed and R has moved to the next line. My R code is something like this:
write_fst(dt, "testfile.fst") a <- 1L # Just to see when fst finishes writing rm(dt); gc() # This is where R hangs for a really long time, sometimes up to 30min
I'll test on my own server later to see if this issue occurs due to Google Cloud or not. Will report back with my findings.
— You are receiving this because you are subscribed to this thread. Reply to this email directly, view it on GitHub https://github.com/fstpackage/fst/issues/228?email_source=notifications&email_token=ABCJ6JLQFO4MZI3UJPKNM2TQ3VII3A5CNFSM4KBWNCCKYY3PNVWWK3TUL52HS4DFVREXG43VMVBW63LNMVXHJKTDN5WW2ZLOORPWSZGOEH5SGCI#issuecomment-570106633, or unsubscribe https://github.com/notifications/unsubscribe-auth/ABCJ6JKTM7EKUFVUSZQAVMLQ3VII3ANCNFSM4KBWNCCA .
@xiaodaigh No I don't. All my files are >95% numeric, with perhaps a few columns of strings. How did you overcome the issue back then when you encountered?
R's gc()? Interesting if the issue does lie there as I've been doing rm(xxx); gc() all this while with no issues whenever there are large objects that are no longer needed. And yes, I've been sticking with R 3.3.3 because it's worked well for me all this while.
I can't overcome it. See if you can start R with gc disabled. You have 900G of RAM so I would check if there are settings limiting your RAM usage. Cos disabling auto gc should do the trick already.
How big were your files when you encountered the hanging issues after write_fst?
About 200GB on a machine with 32GB ram
Hi @winston-p and @xiaodaigh,
yes, the garbage collection can take it's time, although 30 mins seems extremely long. For numerical vectors, the garbage collection should be fast (sub-milliseconds), as a single memory block is allocated for the vector. And freeing memory should not really depend on the size of the vector.
As @xiaodaigh mentions, for character vectors, garbage collection can be relatively slow, especially for vectors with many unique strings, as each string is deleted separately (so that might scale with the size of the vector).
Method write_fst() only allocates small (unmanaged) buffers to process and compress data, but there are no significant R memory allocations done during the write. Therefore, the bulk of garbage collection should consist of deleting the table itself.
Do you see the same delay if you skip the write_fst() line? So, just rm() the table but not write it to disk?
Are you sure the system is not swapping memory during the writes?
thanks
Do you see the same delay if you skip the write_fst() line? So, just rm() the table but not write it to disk?
@MarcusKlik Without write_fst, rm(); gc() has no problem whatsoever. Also, it's not just rm or gc. it happens even with Sys.sleep and stop. As I test further, I'm quite convinced of 4 things:
- The issue has to do with file size: In my workflow, I was writing 50 files of ~3GB each with a lot of processing in-between each
write_fst, no problem. This only started when I was writing files of 20GB and bigger. - It is not due to the level of compression: I tested with 0, 50 and 90, all led to the same issue.
-
read_fstdoes not lead to any hanging; onlywrite_fstdoes. - It has something to do with CPU: In one of my tests, I used only 1 core to run this:
write_fst(dt) # 20GB file
a <- 1L
Sys.sleep(10) # Using stop() leads to hanging too
After writing, R finished a <- 1L very quickly but it hung at Sys.sleep. I noticed then that CPU was at 100% for a long time during the sleep. There was no other app running, just Debian and R. Only after more than 5min (I didn't time it exactly), CPU went back to normal and Sys.sleep completed.
The above summarises the results of my testing. It'd be great if this can be solved as fst truly shines when the files are large.
Hi @winston-p, thanks for testing the issue further.
The strange thing is that fst does not keep any open connections or any threads running after write_fst() returns control to R, so I'm a bit puzzled as to what could be the cause of the observed delay. Just to check that the problem is specific to fst, could you try and run the following code to see if you have the effect?
# generate a 1 GB raw vector
x <- serialize(sample(1:100, 2^28, replace = TRUE), NULL)
# open binary file
tmp_file <- tempfile(fileext = ".bin")
zz <- file(tmp_file, "wb")
# write ~ 27 GB to file
for (block in 0:25) {
writeBin(x, zz)
}
close(zz)
# any delay here?
Sys.sleep(0.01)
thanks!
You're most welcome, @MarcusKlik ! I am really keen to resolve this issue too as it's important for my work.
To simulate the same conditions as when I hit into the issue, I ran your code (with some minor tweaks) on a Google Cloud VM that I cloned from the original one, but this time with 1-core, 60GB ram and a persistent SSD attached.
Summary: there was no delay whatsoever after the write. I went ahead and added a rm(zz); gc() and still there was no delay. One thing I did notice was that, despite there being no hanging, the CPU stayed at 100% for a while after the write and even after R finished running the code.
The code and the associated timings for each crucial step are below:
Timings:
tm1: 2020-01-04 03:32:47
tm2: 2020-01-04 03:37:17
tm3: 2020-01-04 03:37:17
tm4: 2020-01-04 03:37:17
# generate a 1 GB raw vector
x <- serialize(sample(1:100, 2^28, replace = TRUE), NULL)
# open binary file
tmp_file <- tempfile(tmpdir = "/mnt/drive1/TEST", fileext = ".bin")
zz <- file(tmp_file, "wb")
tm1 <- Sys.time()
# write ~ 27 GB to file
for (block in 0:25) {
writeBin(x, zz)
}
close(zz)
tm2 <- Sys.time()
# any delay here?
Sys.sleep(0.01)
tm3 <- Sys.time()
rm(zz); gc()
tm4 <- Sys.time()
cat("tm1:", as.character(tm1), "\n")
cat("tm2:", as.character(tm2), "\n")
cat("tm3:", as.character(tm3), "\n")
cat("tm4:", as.character(tm4), "\n")
I will keep this test VM around for any further tests as and when you.
Sorry, I clicked the wrong button as I was typing the above comment. Re-opening issue.
Hi @winston-p, thanks for testing and cloning the test VM!
So with the above (non-fst) code, there is no delay bit still significant CPU activity, that's still very strange and indicates that garbage collection of large memory blocks takes a lot of CPU power.
Your test-setup includes a persistent SSD that probably has more efficient IO, do you still have the hanging problem with write_fst() on this setup (of just the 100% CPU activity) ?
The two major differences of fst as compared to most other R packages is the high IO speed and the use of OpenMP multithreading.
The first point could strain OS buffering and perhaps, depending on the actual system setup, lead to delays if the IO writes are kept in memory and offloaded to disk later (perhaps a Google Cloud optimization?).
On the second point; OpenMP threads are guaranteed to finish before returning to R, so no delay is expected there. But if somehow OpenMP multithreading introduces a delay after the code has finish, I would expect data.table to exhibit the same effects (data.table uses OpenMP in a similar manner). For example, when writing a large csv with fwrite(). Do you see any delays with fwrite()?
Sorry for the wait, @MarcusKlik - I was rather busy last few weeks.
Your test-setup includes a persistent SSD that probably has more efficient IO, do you still have the hanging problem with write_fst() on this setup (of just the 100% CPU activity) ?
Actually, I added a persistent SSD in the test setup because that's exactly the way it has always been set-up in my main VM, which hit into the write_fst hanging issue in the first place. So the persistent SSD is not the issue.
The first point could strain OS buffering and perhaps, depending on the actual system setup, lead to delays if the IO writes are kept in memory and offloaded to disk later (perhaps a Google Cloud optimization?).
I'm not sure about specific GCE I/O optimizations but my tests with fwrite() did not produce any delays. I ran the code below twice, first with a 1-vCPU server and the second time with a 6-vCPU server, both with 60GB RAM (NOTE: it takes a long time to write the 32-GB CSV file, in case you wish to replicate it).
From the timings below, fwrite does not have the hanging issue I encountered with write_fst().
Kindly let me know what else I can test for you. Thanks!
Timings for 1 vCPU were:
tm1: 2020-01-18 18:03:48
tm2: 2020-01-18 18:18:58
tm3: 2020-01-18 18:18:58
tm4: 2020-01-18 18:18:59
Timings for 6 vCPUs were:
tm1: 2020-01-18 18:30:04
tm2: 2020-01-18 18:38:23
tm3: 2020-01-18 18:38:23
tm4: 2020-01-18 18:38:25
library(data.table) # version 1.10.4
# From: https://www.r-bloggers.com/a-quick-way-to-do-row-repeat-and-col-repeat-rep-row-rep-col/
rep.col<-function(x, n){
matrix(rep(x, each = n), ncol = n, byrow = TRUE)
}
# generate a 32GB DT
nofrow <- 2e6L
nofcol <- 1000L
dt <- as.data.table(rep.col(rnorm(nofrow), nofcol))
# write dt to disk
tm1 <- Sys.time()
tmp_file <- "/mnt/drive1/TEST/dt.csv"
fwrite(dt, tmp_file)
tm2 <- Sys.time()
# any delay here?
Sys.sleep(0.01)
tm3 <- Sys.time()
rm(dt); gc()
tm4 <- Sys.time()
cat("tm1:", as.character(tm1), "\n")
cat("tm2:", as.character(tm2), "\n")
cat("tm3:", as.character(tm3), "\n")
cat("tm4:", as.character(tm4), "\n")
> version
platform x86_64-pc-linux-gnu
arch x86_64
os linux-gnu
system x86_64, linux-gnu
status
major 3
minor 3.3
year 2017
month 03
day 06
svn rev 72310
language R
version.string R version 3.3.3 (2017-03-06)
nickname Another Canoe
Hi @winston-p thanks for testing further!
I'm very puzzled by this delay that you see. From your tests on single core machines, I gather that OpenMP is probably not the issue. On a single core, fst performs all operations on the main thread, so there can't be any (thread-) synchronization problems.
Then you see the delay appearing after the file size exceeds a certain boundary (20 GB) of file size. That's strange because fst doesn't do any buffering except for the blocks that are being written to disk at that time. In other words, memory requirements when writing the start of the file are basically identical to requirements when writing the last bits of the file (so why the 20 GB boundary?).
At the very end of the write operation, fst does jump back to the beginning of the file to (over-)write some metadata in the file header. Normally, suck a seek operation will take a few milliseconds, but perhaps this jump (over > 20 GB) takes a long time on Google Cloud persistent disks?
(does your setup include this type of disk?)
Finally, if you write >20 GB files to disk from datasets that have fewer columns but more rows, is there a delay in those cases (so equal file size but smaller column / row ratio)
Thanks for all your help getting to the bottom of this!
Hi @MarcusKlik , I believe I've found the source of the issue!
After a convoluted round of testing (which I shan't elaborate), I discovered the culprit to be an older version of Rcpp (v0.12.10). After updating to the latest v1.0.3, write_fst no longer hangs.
Sorry for all the confusion and thanks for your patience throughout these few weeks!
Hi @winston-p, great find!
That figures, as Rcpp is in between the calls from R to fstlib and manages some of the memory...
Wow, I'm glad you found the solution, this was a tough nut to crack!
Ok, so we should put a lower bound on the Rcpp version?
Ha @xiaodaigh, we might but on the other hand there are no problems for files smaller than ~20 GB. So putting a lower limit would limit usage to some users that aren't experiencing any problems at the moment.
Perhaps, if we can confirm the delays on other platforms as well (and pinpoint the exact Rcpp version that provides a fix for that), we can emit a warning at startup?
@xiaodaigh and @MarcusKlik , perhaps we can leave it as it is unless someone else faces the same issue in future and found that indeed Rcpp is the solution? My case is just but one incident.
My 2 cents for your consideration.
Yes, agreed, if this issue resurfaces, we can investigate and put the lower boundary on Rcpp if needed.
Thanks for all the testing and suggestions!