Mean Diffusivity turns out low in CSF when reconstructing using ReconstDtiFlow
Description
MD appears to be low in CSF voxels when computing it via DIPY's ReconstDtiFlow for reconstructing the diffusion tensor.
This behavior is somewhat unexpected, especially due to the same voxels being given high MD values when computed via DIPY's ReconstDkiFlow for reconstructing the tensor using the kurtosis model.
My data is multi-shelled, with the following shells: 4 b-value shells (0, 1000, 2000, and 4000) with 6 images measured at b=0 and 60 directions for each of the rest (1000, 2000, and 4000).
Tensors were fitted using the Weighted Least Squares (WLS) method as I think is required due to the high b values present in this scheme.
Attached is a single-subject MD map, calculated via the Tensor and Kurtosis models:
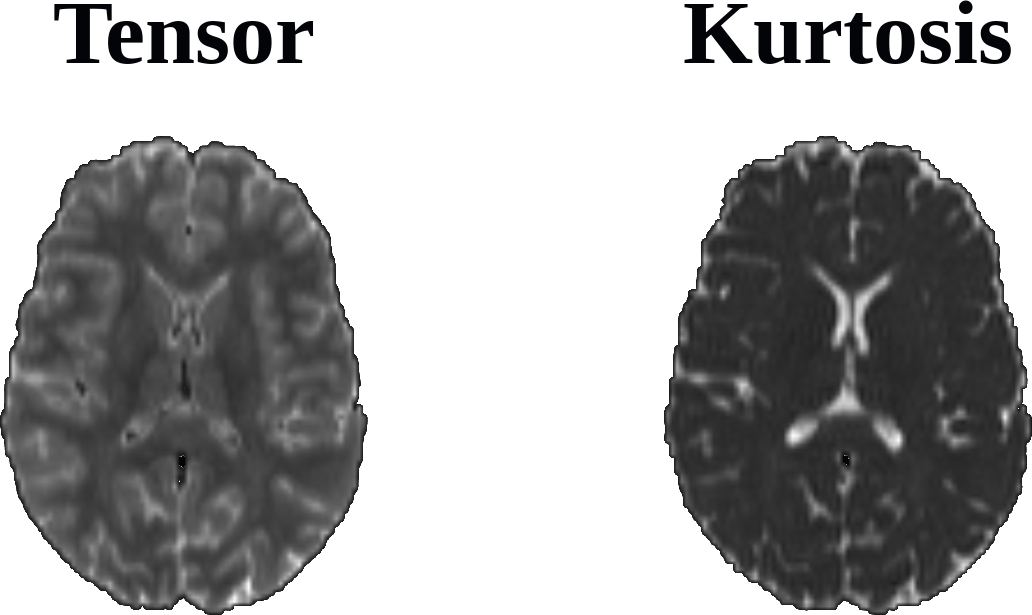
I was wondering whether this issue looks familiar to anyone that may help me shed some light on this matter as it got me very confused :smiling_face_with_tear:
Way to reproduce
- [x] Relevant images (if any)
- [x] Operating system and version Linux-5.13.0-44-generic-x86_64-with-glibc2.31
- [x] Python version Python 3.9.13
- [x] dipy version 1.5.0
- [x] dependency version (numpy, scipy, nibabel, h5py, cvxpy, fury)
- NumPy 1.22.3
- SciPy 1.8.0
- Nibabel 3.2.2
- H5py 3.6.0
Any update on this issue ?
Sorry I missed this question when it was originally posted. This is not surprising and arises from the inclusion of high b-values in the DTI fit. When you include these measurements, you are including data that is very close to the noise floor and therefore, the MD value is biased downward. The following image from Desantis et al. shows why this happens, and how DKI mitigates it:
The dashed line represents the physiological reaIity, but because noise in MRI data is non-negative (because it's a sum of squares, see this paper), the data close to the noise floor of the measurement (i.e., where the signal is so close to 0 that it cannot be distinguished from the noise) is always positively biased. This means that if you were to fit a straight line (which is what DTI does) to those points going out all the way to b=5,000, you would get a line with a slope that is shallower than it should be. DKI explicitly avoids this issue by fitting a quadratic function to this line. One sanity check you should always do is to check whether the CSF in the ventricles has a MD of approximately 3 mm^2/s, which is what you expect for water at body temp. If you find that your MD there is lower than expected, you might consider whether the SNR of your measurements (especially at higher b-values) is too low to use with this model. A common solution is to fit DKI only to b<2,000 measurements, or to use DKI.
Thanks a lot for the quick response and the insights. For future reference, here is a comparison of different MD estimations on multi-shell data (b=1000,2000,3000,5000) with each shell estimated separately (first row), cumulated (second row) and the two solutions advised above (third row). Both of which correctly estimate the diffusivity to be around 0.003 mm^2/s. I have added a third option in the last row, which is a rough estimate based on the combination of all shells, evaluated separately. This last option is just to spark a discussion, as it has the look I expected a multi-shell estimate of MD to have, with a clear distinction of the CSF, grey matter, white matter and even the internal capsule. I will look further into it to make sure that it makes sense theoretically and is robust to different acquisitions.
This is great! Textbook image right there.
I am not sure that I understand how the proposed image is generated. Do you calculate DTI in each shell and then combine the estimates somehow? Averaging with equal weighting for each shell??