phy
 phy copied to clipboard
phy copied to clipboard
Unit wrongly assigned to channel
I have found sometimes that a cluster is assigned with best channel a channel that doesn't have the spike. For example in this case, I have a cluster assigned with best channel at channel 0, but the true unit is at channel 11.

This is more of a problem when the unit is in a channel farther away, so it doesn't even appear in the waveform view, and my only way to see it, is through the traceview
What could be the reason? This dataset was sorted with Kilosort 2, I'm not sure if this is a phy problem or KS2 problem.
Maybe this is related to what has also been observed when data are sorted with SpyKING CIRCUS #1014?
I'm currently refactoring the logic assigning best channels to clusters (dev branch of phylib), it would be helpful if you could try it out and, if the problem persists, send me your dataset with a description of what's wrong so that I can debug properly. Thanks.
I just did
pip install --upgrade git+https://github.com/cortex-lab/phy
pip install --upgrade git+https://github.com/cortex-lab/phylib@dev
rm -rf .phy
and now I get this error
13:15:29.724 [I] model:276 File format is ks2.
13:15:29.760 [W] model:596 Unreferenced clusters found in spike_clusters (generally not a problem)
13:15:29.772 [E] __init__:62 An error has occurred (IndexError): index 163 is out of bounds for axis 0 with size 32
Traceback (most recent call last):
File "/home/ariel/virtualenvs/phy2/.phy2/bin/phy", line 11, in <module>
load_entry_point('phy==2.0b2.dev0', 'console_scripts', 'phy')()
File "/home/ariel/virtualenvs/phy2/.phy2/lib/python3.6/site-packages/click/core.py", line 829, in __call__
return self.main(*args, **kwargs)
File "/home/ariel/virtualenvs/phy2/.phy2/lib/python3.6/site-packages/click/core.py", line 782, in main
rv = self.invoke(ctx)
File "/home/ariel/virtualenvs/phy2/.phy2/lib/python3.6/site-packages/click/core.py", line 1259, in invoke
return _process_result(sub_ctx.command.invoke(sub_ctx))
File "/home/ariel/virtualenvs/phy2/.phy2/lib/python3.6/site-packages/click/core.py", line 1066, in invoke
return ctx.invoke(self.callback, **ctx.params)
File "/home/ariel/virtualenvs/phy2/.phy2/lib/python3.6/site-packages/click/core.py", line 610, in invoke
return callback(*args, **kwargs)
File "/home/ariel/virtualenvs/phy2/.phy2/lib/python3.6/site-packages/click/decorators.py", line 21, in new_func
return f(get_current_context(), *args, **kwargs)
File "/home/ariel/virtualenvs/phy2/.phy2/lib/python3.6/site-packages/phy/apps/__init__.py", line 159, in cli_template_gui
template_gui(params_path, **kwargs)
File "/home/ariel/virtualenvs/phy2/.phy2/lib/python3.6/site-packages/phy/apps/template/gui.py", line 209, in template_gui
model = load_model(params_path)
File "/home/ariel/virtualenvs/phy2/.phy2/lib/python3.6/site-packages/phylib/io/model.py", line 1495, in load_model
return TemplateModel(**get_template_params(params_path))
File "/home/ariel/virtualenvs/phy2/.phy2/lib/python3.6/site-packages/phylib/io/model.py", line 263, in __init__
self._load_data()
File "/home/ariel/virtualenvs/phy2/.phy2/lib/python3.6/site-packages/phylib/io/model.py", line 356, in _load_data
self.sparse_templates = self._load_templates()
File "/home/ariel/virtualenvs/phy2/.phy2/lib/python3.6/site-packages/phylib/io/model.py", line 768, in _load_templates
mat = self.wmi[np.ix_(channel_ids, channel_ids)]
IndexError: index 163 is out of bounds for axis 0 with size 32
QWidget: Must construct a QApplication before a QWidget
Aborted (core dumped)
I can go back with
pip install --upgrade git+https://github.com/cortex-lab/phylib
rm -rf .phy
and phy lunchs again
could you send me your dataset (no need for the raw data)?
https://www.dropbox.com/s/mh9nd4qyd7wrbun/data1.zip?dl=0
Thanks. I'm unable to reproduce the issue on phylib@dev. Could you double check that, if you export this zip file in a fresh folder, and do phy template-gui params.py --debug in it, you continue to see the error message on your computer?
I just did a fresh install of phy and phylib@dev
pip install --upgrade git+https://github.com/cortex-lab/phy
pip install --upgrade git+https://github.com/cortex-lab/phylib@dev
pip install --upgrade qtwebengine (otherwise there is an error: ModuleNotFoundError: No module named 'PyQt5.QtWebEngineWidgets')
phy template-gui params.py --debug
12:12:30.583 [D] __init__:68 Start capturing exceptions.
12:12:30.583 [W] model:1469 File continuous.dat does not exist.
12:12:30.584 [I] model:276 File format is ks2.
12:12:30.609 [D] model:589 Loading spike clusters.
12:12:30.617 [W] model:596 Unreferenced clusters found in spike_clusters (generally not a problem)
12:12:30.626 [D] model:518 No channel shank file found.
12:12:30.626 [D] model:819 Loading the whitening matrix.
12:12:30.627 [D] model:826 Loading the inverse of the whitening matrix.
12:12:30.627 [D] model:671 Loading templates.
12:12:30.628 [D] model:706 Open KS2 template waveforms.
12:12:30.629 [D] model:719 Templates are sparse.
12:12:30.630 [E] __init__:62 An error has occurred (IndexError): index 163 is out of bounds for axis 0 with size 32
Traceback (most recent call last):
File "/home/ariel/virtualenvs/phyt/.phy2/bin/phy", line 11, in <module>
load_entry_point('phy==2.0b2.dev0', 'console_scripts', 'phy')()
File "/home/ariel/virtualenvs/phyt/.phy2/lib/python3.6/site-packages/click/core.py", line 829, in __call__
return self.main(*args, **kwargs)
File "/home/ariel/virtualenvs/phyt/.phy2/lib/python3.6/site-packages/click/core.py", line 782, in main
rv = self.invoke(ctx)
File "/home/ariel/virtualenvs/phyt/.phy2/lib/python3.6/site-packages/click/core.py", line 1259, in invoke
return _process_result(sub_ctx.command.invoke(sub_ctx))
File "/home/ariel/virtualenvs/phyt/.phy2/lib/python3.6/site-packages/click/core.py", line 1066, in invoke
return ctx.invoke(self.callback, **ctx.params)
File "/home/ariel/virtualenvs/phyt/.phy2/lib/python3.6/site-packages/click/core.py", line 610, in invoke
return callback(*args, **kwargs)
File "/home/ariel/virtualenvs/phyt/.phy2/lib/python3.6/site-packages/click/decorators.py", line 21, in new_func
return f(get_current_context(), *args, **kwargs)
File "/home/ariel/virtualenvs/phyt/.phy2/lib/python3.6/site-packages/phy/apps/__init__.py", line 159, in cli_template_gui
template_gui(params_path, **kwargs)
File "/home/ariel/virtualenvs/phyt/.phy2/lib/python3.6/site-packages/phy/apps/template/gui.py", line 209, in template_gui
model = load_model(params_path)
File "/home/ariel/virtualenvs/phyt/.phy2/lib/python3.6/site-packages/phylib/io/model.py", line 1495, in load_model
return TemplateModel(**get_template_params(params_path))
File "/home/ariel/virtualenvs/phyt/.phy2/lib/python3.6/site-packages/phylib/io/model.py", line 263, in __init__
self._load_data()
File "/home/ariel/virtualenvs/phyt/.phy2/lib/python3.6/site-packages/phylib/io/model.py", line 356, in _load_data
self.sparse_templates = self._load_templates()
File "/home/ariel/virtualenvs/phyt/.phy2/lib/python3.6/site-packages/phylib/io/model.py", line 768, in _load_templates
mat = self.wmi[np.ix_(channel_ids, channel_ids)]
IndexError: index 163 is out of bounds for axis 0 with size 32
QWidget: Must construct a QApplication before a QWidget
Aborted (core dumped)
I tried a different dataset sorted with kilosort2 and same problem I tried with a dataset sorted with spyking-circus and it lunchs. Not really sure what could be the problem.
I believe I have similar experience.
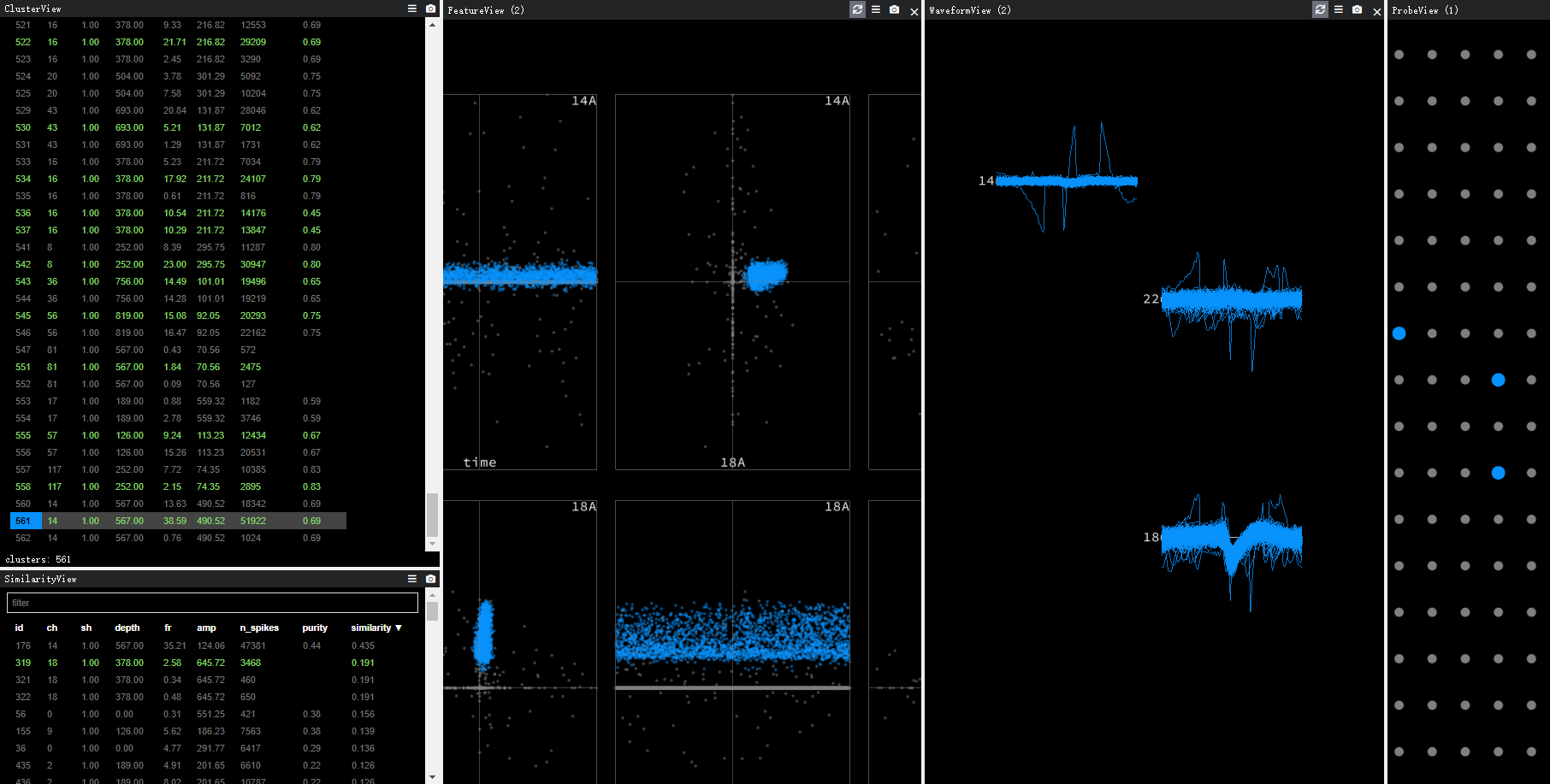
In this picture, true unit is at ch 18 ,while ch 14 is chosen.
Seems that the algorithm is choosing the weakest amplitude rather than the largest.
This could be problematic if one is concerned about the precise location of neurons recorded.
Hi @rossant could you please suggest what we should do to fix the channel assignment in a quick way? Or which module of phy should we look at?
I'm using phy TemplateGUI v2.0b2.dev, and phylib v2.3a0. I believe this is already the most recent dev version.
Update:
I'm using Spyking-Circus, and I found after setting the "sparse_export" option set to False, then Phy could find the correct channel now.