some contourf levels not drawn in Orthographic projection (are drawn by contour)
Description
I'm trying to draw spherical harmonic contours on an orthographic projection of a sphere. I see the correct contours when calling contour(), but many are not filled by contourf(). I think this is a bug, perhaps related to Issue 1024 (but I couldn't view their example figures).
Code to reproduce
import matplotlib.pyplot as plt
import numpy as np
import scipy.special as sp
import cartopy
import cartopy.crs as ccrs
# Compute spherical harmonic pattern
l=4; m=3
lon = np.linspace(0,2*np.pi,200)
lat = np.linspace(-np.pi/2,np.pi/2,500)
colat = lat+np.pi/2
d = np.zeros((len(lon),len(colat)),dtype = np.complex64)
meshed_grid = np.meshgrid(lon, lat)
lat_grid = meshed_grid[1]
lon_grid = meshed_grid[0]
for j, yy in enumerate(colat):
for i, xx in enumerate(lon):
d[i,j] = sp.sph_harm(m,l,xx,yy)
drm = np.transpose(np.real(d))
#Plot example with contour() and contourf()
fig, (ax1,ax2) = plt.subplots(1, 2, subplot_kw={'projection': ccrs.Orthographic(0, 30)})
#contour() works as expected
ax1.contour(lon*180/np.pi,lat*180/np.pi,drm,51,
transform=ccrs.PlateCarree(),cmap='seismic')
ax1.relim()
ax1.autoscale_view()
#contourf() doesn't
ax2.contourf(lon*180/np.pi,lat*180/np.pi,drm,51,
transform=ccrs.PlateCarree(),cmap='seismic')
ax2.relim()
ax2.autoscale_view()
plt.tight_layout()
plt.savefig('sphericalcontours.png')

Traceback
No error reported.
Full environment definition
Operating system
OS X 10.10.5
Cartopy version
0.16.0
conda list
# packages in environment at /Users/keatonb/anaconda:
#
# Name Version Build Channel
_license 1.1 py27_0 <unknown>
abstract-rendering 0.5.1 np19py27_0 <unknown>
ads 0.12.3 <pip>
alabaster 0.7.6 py27_0
anaconda 2.3.0 np19py27_0 <unknown>
anaconda-client 1.5.1 py27_0
anaconda-navigator 1.3.1 py27_0
aplpy 1.1.1 py27_0 astropy
appnope 0.1.0 py27_0
appscript 1.0.1 py27_0 <unknown>
argcomplete 0.8.9 py27_0 <unknown>
asdf 1.2.1 py27_0 astropy
astro-gala 0.1.3 py27_0 astropy
astroid 1.4.7 py27_0
astroml 0.3 py27_0 astropy
astroML 0.3 <pip>
astroplan 0.2 py27_0 astropy
astropy 1.0.4 <pip>
astropy 1.3.3 <pip>
astropy 1.1.1 <pip>
astropy 2.0.3 py27_0 conda-forge
astropy-helpers 1.1.1 0 http://eupsforge.net/conda/dev
astroquery 0.3.7 py27_2 astropy
astroscrappy 1.0.5 np111py27_0 astropy
babel 2.1.1 py27_0
backports 1.0 py27_0
backports.functools_lru_cache 1.5 py27_1
backports_abc 0.4 py27_0
basemap 1.1.0 py27_3 conda-forge
bcolz 0.9.0 np19py27_0 <unknown>
beautiful-soup 4.3.2 py27_0 <unknown>
beautifulsoup4 4.5.1 py27_0
binstar 0.11.0 py27_0 <unknown>
bitarray 0.8.1 py27_0 <unknown>
blaze-core 0.8.0 np19py27_0 <unknown>
bleach 1.5.0 py27_0 conda-forge
blinker 1.4 <pip>
blz 0.6.2 np19py27_1 <unknown>
bokeh 0.12.16 py27_0
boto 2.38.0 py27_0 <unknown>
bottleneck 1.0.0 np110py27_0
brewer2mpl 1.4.1 <pip>
ca-certificates 2018.03.07 0
cartopy 0.16.0 py27he7b4726_0
ccdproc 1.2.0 py27_0 astropy
cdecimal 2.3 py27_0 <unknown>
certifi 2018.4.16 py27_0
cffi 1.9.1 py27_0
chardet 3.0.4 py27h2842e91_1
chardet 2.3.0 <pip>
chest 0.2.3 py27_0
clangdev 5.0.0 default_0 conda-forge
click 6.6 py27_0
click-plugins 1.0.3 py27_0
cligj 0.4.0 py27_0
cloudpickle 0.2.2 py27_0
cluster-lensing 0.1.2 py27_0 astropy
clyent 0.3.4 py27_0 <unknown>
colorama 0.3.3 py27_0 <unknown>
conda 4.5.4 py27_0
conda-build 1.14.1 py27_0 <unknown>
conda-env 2.6.0 h36134e3_0
configobj 5.0.6 py27_0 <unknown>
configparser 3.5.0b2 py27_1
corner 2.0.1 <pip>
coverage 3.7.1 <pip>
cryptography 1.7.1 py27_0
curl 7.49.0 1
cycler 0.10.0 py27_0
cython 0.24 py27_0
cytoolz 0.7.3 py27_0 <unknown>
d2to1 0.2.12.post1 0 http://eupsforge.net/conda/dev
dask 0.10.0 <pip>
dask 0.16.0 py27h0cfea73_0
dask-core 0.16.0 py27h3e6bc05_0
datashape 0.4.5 np19py27_0 <unknown>
decorator 4.0.9 py27_0
decorator 4.0.10 <pip>
dill 0.2.7.1 py27hc0d316d_0
distributed 1.20.2 py27_0
docutils 0.12 py27_0 <unknown>
dynd-python 0.6.5 np19py27_0 <unknown>
emcee 2.2.1 py27_1 astropy
entrypoints 0.2 py27_1
enum34 1.1.6 py27_0
enum34 1.1.2 <pip>
eups 2.0.2 0 http://eupsforge.net/conda/dev
everest-pipeline 2.0.8 <pip>
extinction 0.3.0 np112py27_0 conda-forge
fastcache 1.0.2 py27_0 <unknown>
feedgenerator 1.7 <pip>
flask 0.10.1 py27_1 <unknown>
freetype 2.8 h12048fb_1
funcsigs 0.4 py27_0 <unknown>
funcsigs 1.0.2 <pip>
functools32 3.2.3.2 py27_0
future 0.16.0 py27_0
futures 3.2.0 py27_0 conda-forge
galpy 1.2 <pip>
gammapy 0.4 py27_0 astropy
gatspy 0.3 <pip>
george 0.2.1 <pip>
geos 3.6.2 1 conda-forge
geotiff 1.4.2 h54263a3_0
get_terminal_size 1.0.0 py27_0
gevent 1.0.1 py27_0 <unknown>
gevent-websocket 0.9.3 py27_0 <unknown>
ginga 2.6.1 py27_0 astropy
glue-core 0.12.4 py27_0
glue-vispy-viewers 0.9.2 py27_0
glueviz 0.12.4 0
gPhoton 1.28.2 <pip>
greenlet 0.4.7 py27_0 <unknown>
grin 1.2.1 py27_1 <unknown>
gwcs 0.7 py27_0 astropy
h5py 2.7.0 np112py27_0 conda-forge
halotools 0.4 py27_0 astropy
hdf4 4.2.13 0 conda-forge
hdf5 1.8.17 1
healpy 1.8.6 <pip>
heapdict 1.0.0 py27_1
html5lib 0.9999999 py27_0 conda-forge
httpretty 0.8.14 py27h7612536_1
httpretty 0.8.10 <pip>
icu 54.1 0
idna 2.6 py27hedea723_1
imageio 2.2.0 py27_0 conda-forge
imageio 1.5 <pip>
imagesize 0.7.1 py27_0
imexam 0.7.0 py27_0 astropy
ipaddress 1.0.7 py27_0 <unknown>
ipykernel 4.6.1 py27_0
ipython 5.3.0 py27_0
ipython-notebook 4.0.4 py27_0
ipython-qtconsole 4.0.1 py27_0
ipython_genutils 0.1.0 py27_0
ipywidgets 6.0.0 py27_0
itsdangerous 0.24 py27_0 <unknown>
jbig 2.1 0
jdcal 1.0 py27_0 <unknown>
jedi 0.9.0 py27_0
jinja2 2.8 py27_0
jpeg 9b 2 conda-forge
jsonschema 2.4.0 py27_0 <unknown>
jupyter_client 4.2.2 py27_0
jupyter_core 4.1.0 py27_0
k2flix 1.1.dev0 <pip>
K2fov 6.2.0 <pip>
kealib 1.4.6 0
keyring 9.0 py27_0 astropy
kiwisolver 1.0.1 py27h9856860_0
launcher 1.0.0 3 <unknown>
lazy-object-proxy 1.2.1 py27_0
legacy_configs 1.0.0 py27_0 http://eupsforge.net/conda/dev
libcxx 5.0.0 0 conda-forge
libdynd 0.6.5 0 <unknown>
libgdal 1.11.2 1
libgfortran 3.0.1 h93005f0_2
libnetcdf 4.4.1 0
libopenblas 0.2.20 h6c53463_3
libpng 1.6.34 he12f830_0
libsodium 0.4.5 0 <unknown>
libtiff 4.0.9 hcb84e12_1
libxml2 2.9.2 0 <unknown>
libxslt 1.1.29 4 conda-forge
llvmdev 5.0.0 default_0 conda-forge
llvmlite 0.9.0 py27_0
lmfit 0.9.5 py27_0 astropy
locket 0.2.0 py27_1
lsst-afw w.2016.15 0 http://eupsforge.net/conda/dev
lsst-astrometry-net 0.50.2.post6 0 http://eupsforge.net/conda/dev
lsst-astrometry-net-data 0.10.0.post68 0 http://eupsforge.net/conda/dev
lsst-base w.2016.15 0 http://eupsforge.net/conda/dev
lsst-boost 1.59.5 0 http://eupsforge.net/conda/dev
lsst-cfitsio 3360.4 0 http://eupsforge.net/conda/dev
lsst-coadd-chisquared w.2016.15 0 http://eupsforge.net/conda/dev
lsst-coadd-utils w.2016.15 0 http://eupsforge.net/conda/dev
lsst-daf-base w.2016.15 0 http://eupsforge.net/conda/dev
lsst-daf-butlerutils w.2016.15 0 http://eupsforge.net/conda/dev
lsst-daf-persistence w.2016.15 0 http://eupsforge.net/conda/dev
lsst-doxygen 1.8.5.1 0 http://eupsforge.net/conda/dev
lsst-eigen 3.2.5.1 70497dd_0 http://eupsforge.net/conda/dev
lsst-esutil 0.5.3 0 http://eupsforge.net/conda/dev
lsst-fftw 3.3.4.2 0 http://eupsforge.net/conda/dev
lsst-freetds 0.91.112.2 0 http://eupsforge.net/conda/dev
lsst-galsim 1.3.2 0 http://eupsforge.net/conda/dev
lsst-geom w.2016.15 0 http://eupsforge.net/conda/dev
lsst-gsl 1.16.3 0 http://eupsforge.net/conda/dev
lsst-healpy 1.8.1.2.post1 d4cac0b_0 http://eupsforge.net/conda/dev
lsst-ip-diffim w.2016.15 0 http://eupsforge.net/conda/dev
lsst-ip-isr w.2016.15 0 http://eupsforge.net/conda/dev
lsst-mariadbclient 0.10.1.11.2 0 http://eupsforge.net/conda/dev
lsst-meas-algorithms w.2016.15 0 http://eupsforge.net/conda/dev
lsst-meas-astrom w.2016.15 0 http://eupsforge.net/conda/dev
lsst-meas-base w.2016.15 0 http://eupsforge.net/conda/dev
lsst-meas-deblender w.2016.15 0 http://eupsforge.net/conda/dev
lsst-minuit2 5.28.00.2.1 dae2fb7_0 http://eupsforge.net/conda/dev
lsst-mysqlclient 5.1.73.2 2 http://eupsforge.net/conda/dev
lsst-mysqlpython 1.2.3.1.1.post6 120147b_0 http://eupsforge.net/conda/dev
lsst-ndarray w.2016.15 0 http://eupsforge.net/conda/dev
lsst-obs-lsstsim w.2016.15 0 http://eupsforge.net/conda/dev
lsst-obs-test w.2016.15 0 http://eupsforge.net/conda/dev
lsst-palpy 1.7.0.1.1 733f88b_0 http://eupsforge.net/conda/dev
lsst-pex-config w.2016.15 0 http://eupsforge.net/conda/dev
lsst-pex-exceptions w.2016.15 0 http://eupsforge.net/conda/dev
lsst-pex-logging w.2016.15 0 http://eupsforge.net/conda/dev
lsst-pex-policy w.2016.15 0 http://eupsforge.net/conda/dev
lsst-pipe-base w.2016.15 0 http://eupsforge.net/conda/dev
lsst-pipe-tasks w.2016.15 0 http://eupsforge.net/conda/dev
lsst-product-configs 1.0.1925 0 http://eupsforge.net/conda/dev
lsst-pyephem 3.7.5.1.1 6 http://eupsforge.net/conda/dev
lsst-pyfits 3.4.0.post2 0 http://eupsforge.net/conda/dev
lsst-pykg-config 1.2.0.post5 3 http://eupsforge.net/conda/dev
lsst-pymssql 2.1.1.post3 0 http://eupsforge.net/conda/dev
lsst-python-d2to1 0.2.12.1 0 http://eupsforge.net/conda/dev
lsst-scons 2.3.5 0 http://eupsforge.net/conda/dev
lsst-sconsutils w.2016.15 0 http://eupsforge.net/conda/dev
lsst-sims master_gac1855abae.0058 ac1855abae_0 http://eupsforge.net/conda/dev
lsst-sims-catalogs-generation sims_2.2.4.post2 0 http://eupsforge.net/conda/dev
lsst-sims-catalogs-measures sims_2.2.4.post2 0 http://eupsforge.net/conda/dev
lsst-sims-catutils sims_2.2.4.post2 0 http://eupsforge.net/conda/dev
lsst-sims-coordutils sims_2.2.4.post2 0 http://eupsforge.net/conda/dev
lsst-sims-data sims_2.2.4 0 http://eupsforge.net/conda/dev
lsst-sims-dustmaps 0.10.1.1 57fdb73_2 http://eupsforge.net/conda/dev
lsst-sims-galsiminterface sims_2.2.4.post2 0 http://eupsforge.net/conda/dev
lsst-sims-maf sims_2.2.4maf.post1 0 http://eupsforge.net/conda/dev
lsst-sims-maps sims_2.2.4 0 http://eupsforge.net/conda/dev
lsst-sims-photutils sims_2.2.4.post2 0 http://eupsforge.net/conda/dev
lsst-sims-sed-library 2016.01.26 0 http://eupsforge.net/conda/dev
lsst-sims-utils sims_2.2.4.post2 0 http://eupsforge.net/conda/dev
lsst-skymap w.2016.15 0 http://eupsforge.net/conda/dev
lsst-skypix w.2016.15 0 http://eupsforge.net/conda/dev
lsst-sncosmo 1.2.0.post1 0 http://eupsforge.net/conda/dev
lsst-stsci-distutils 0.3.7.1.post1 b22a065_0 http://eupsforge.net/conda/dev
lsst-throughputs sims_2.2.4 0 http://eupsforge.net/conda/dev
lsst-tmv 0.72.4 0 http://eupsforge.net/conda/dev
lsst-utils w.2016.15 0 http://eupsforge.net/conda/dev
lsst-wcslib 5.13.1 0 http://eupsforge.net/conda/dev
lxml 4.1.1 py27_0 conda-forge
maltpynt 2.0 py27_0 astropy
Markdown 2.6.5 <pip>
markdown 2.6.9 py27_0 conda-forge
markupsafe 0.23 py27_0 <unknown>
matplotlib 2.2.2 py27ha7267d0_0
matplotlib 2.0.2 <pip>
mistune 0.7.2 py27_1
mkl 11.3.3 0
mock 2.0.0 py27_0 conda-forge
modernize 0.5 <pip>
montage-wrapper 0.9.8 py27_0 astropy
mpld3 0.2 <pip>
mpldatacursor 0.6.2 <pip>
mpmath 0.19 py27_1
msgpack-python 0.4.8 py27h635ded4_0
multipledispatch 0.4.7 py27_0 <unknown>
munch 2.0.4 py27_0
naima 0.8 py27_0 astropy
nbconvert 4.2.0 py27_0
nbformat 4.0.1 py27_0
networkx 1.10 py27_0
networkx 1.11 <pip>
nltk 3.0.3 np19py27_0 <unknown>
node-webkit 0.10.1 0 <unknown>
nomkl 1.0 0
nose 1.3.7 py27_0 <unknown>
notebook 5.0.0 py27_0
numba 0.24.0 np110py27_0
numexpr 2.6.4 py27_nomklh7ee8dc7_0 [nomkl]
numpy 1.12.1 py27_nomkl_0 [nomkl]
numpy 1.12.1 <pip>
numpy 1.14.0 <pip>
numpy 1.11.1 <pip>
numpy 1.10.4 <pip>
numpydoc 0.6.0 py27_0
odo 0.3.2 np19py27_0 <unknown>
olefile 0.45.1 py27_0
omnifit 0.2.1 py27_0 astropy
openpyxl 1.8.5 py27_0 <unknown>
openssl 1.0.2o h26aff7b_0
owslib 0.16.0 py_0 conda-forge
packaging 17.1 py27_0
palpy 1.6.0 <pip>
pandas 0.21.1 py27h1cb45b9_0
pandas-qt 0.1.2 <pip>
partd 0.3.8 py27h7560dbf_0
path.py 8.2 py27_0
pathlib2 2.2.1 py27_0
patsy 0.4.1 py27_0
pbr 3.1.1 py27_0 conda-forge
pcre 8.31 0
pelican 3.6.3 <pip>
pep8 1.6.2 py27_0 <unknown>
pexpect 4.0.1 py27_0
photutils 0.2 <pip>
photutils 0.3 py27_0 astropy
pickleshare 0.5 py27_0
Pillow 3.3.0 <pip>
pillow 4.2.1 py27h2cf1d5f_0
pip 10.0.1 <pip>
pip 9.0.1 py27h1567d89_4
pip 9.0.1 <pip>
plotly 2.0.15 py27h2ba4078_0
ply 3.6 py27_0 <unknown>
probableparsing 0.0.1 <pip>
proj4 4.9.3 5 conda-forge
prompt_toolkit 1.0.14 py27_0
protobuf 3.5.1 py27_3 conda-forge
psutil 3.3.0 py27_0
ptyprocess 0.5 py27_0
py 1.4.27 py27_0 <unknown>
py 1.5.3 <pip>
pyasn1 0.1.9 py27_0
pyaudio 0.2.7 py27_0 <unknown>
pycodestyle 2.3.1 py27_0
pycosat 0.6.3 py27h6c51c7e_0
pycparser 2.14 py27_0 <unknown>
pycrypto 2.6.1 py27_4
pycurl 7.19.5.1 py27_2 <unknown>
pydl 0.5.3 py27_0 astropy
pyephem 3.7.5.3 <pip>
pyepsg 0.3.2 py27_0 conda-forge
pyfits 3.4 <pip>
pyflakes 1.0.0 py27_0
pygments 2.1.3 py27_0
pyketools 3.0b3 <pip>
pylint 1.5.4 py27_1
pymc 2.3.6 py27_1 conda-forge
pyopengl 3.1.1a1 np112py27_0
pyopenssl 16.2.0 py27_0
pyparsing 2.1.1 py27_0
pyparsing 2.2.0 <pip>
pyproj 1.9.5.1 py27_0
pyqt 5.6.0 py27_2
pyqtgraph 0.9.10 <pip>
pyregion 2.0 py27_0 conda-forge
pyshp 1.2.12 py_0 conda-forge
pysocks 1.6.8 py27_0
pysqlite 2.8.3 <pip>
pysyzygy 0.0.1 <pip>
pytables 3.2.0 np19py27_0 <unknown>
pytest 2.9.1 py27_0
pytest-cov 2.1.0 <pip>
pytest-qt 1.2.2 <pip>
python 2.7.12 1
python-cpl 0.7.2 py27_0 astropy
python-crfsuite 0.9.2 py27_0 conda-forge
python-dateutil 2.6.0 <pip>
python-dateutil 2.5.2 py27_0
python.app 1.2 py27_4 <unknown>
pytz 2017.2 <pip>
pytz 2016.3 py27_0
pyvo 0.5.1 <pip>
pywavelets 0.5.2 py27_1 conda-forge
pyyaml 3.11 py27_1 <unknown>
pyzipcode 1.0 <pip>
pyzmq 15.2.0 py27_0
qt 5.6.2 0
qtawesome 0.4.1 py27_0
qtconsole 4.3.0 py27_0
qtpy 1.2.0 py27_0
readline 6.2 2
redis 2.6.9 0 <unknown>
redis-py 2.10.3 py27_0 <unknown>
reproject 0.3.2 py27_0 conda-forge
requests 2.18.4 py27h9b2b37c_1
requests 2.11.0 <pip>
rope 0.9.4 py27_1 <unknown>
ruamel_yaml 0.11.14 py27_0
runipy 0.1.3 py27_0 <unknown>
scandir 1.5 py27_0
scikit-image 0.13.0 py27_1 conda-forge
scikit-image 0.12.3 <pip>
scikit-learn 0.19.1 py27_nomklhd1ada06_0 [nomkl]
scipy 0.19.0 <pip>
scipy 1.0.0 py27_nomklhb534402_0 [nomkl]
scons 2.3.0 py27_0
seaborn 0.7.1 py27_0
setuptools 18.3.2 <pip>
setuptools 38.4.0 py27_0 anaconda
setuptools-git 1.1 1 http://eupsforge.net/conda/dev
shapely 1.6.3 py27_0 conda-forge
simplegeneric 0.8.1 py27_0
singledispatch 3.4.0.3 py27_0
sip 4.18 py27_0
six 1.10.0 py27_0
sncosmo 1.4.0 py27_0 astropy
snowballstemmer 1.2.0 py27_0 <unknown>
sockjs-tornado 1.0.1 py27_0 <unknown>
sortedcontainers 1.5.7 py27h322dbbf_0
spectral-cube 0.4.0 py27_0 astropy
specutils 0.2.2 py27_0 astropy
spherical-geometry 1.0.6 py27_0 astropy
sphinx 1.4.6 py27_0
sphinx_rtd_theme 0.1.7 py27_0 <unknown>
spyder 3.2.8 py27_0
spyder-app 2.3.8 py27_0
sqlalchemy 1.0.12 py27_0
sqlite 3.13.0 0
ssl_match_hostname 3.4.0.2 py27_1
statsmodels 0.8.0 py27_0 conda-forge
stsci.distutils 0.3.7 1 http://eupsforge.net/conda/dev
subprocess32 3.2.7 py27_0
swig 3.0.2 0
sympy 1.0 py27_0
tblib 1.3.2 py27ha684fc4_0
tensorboard 0.4.0rc3 py27_2 conda-forge
tensorflow 1.4.0 py27_0 conda-forge
terminado 0.5 py27_1
titlecase 0.12.0 <pip>
tk 8.5.18 0
toolz 0.8.0 <pip>
toolz 0.8.2 py27h27228c4_0
tornado 4.5.2 py27h29aec9e_0
tqdm 4.14.0 py27_0
traitlets 4.3.2 py27_0
ujson 1.33 py27_0 <unknown>
unicodecsv 0.9.4 py27_0 <unknown>
Unidecode 0.4.19 <pip>
urllib3 1.22 py27hc3787e9_0
usaddress 0.5.10 <pip>
wcsaxes 0.9 py27_0 astropy
wcwidth 0.1.7 py27_0
webencodings 0.5 py27_0 conda-forge
werkzeug 0.12.2 py_1 conda-forge
wheel 0.29.0 py27_0
widgetsnbextension 2.0.0 py27_0
wikipedia 1.4.0 <pip>
wrapt 1.10.8 py27_0
xerces-c 3.1.4 0
xlrd 1.1.0 py27hbd41ed1_1
xlsxwriter 0.7.3 py27_0 <unknown>
xlwings 0.3.5 py27_0 <unknown>
xlwt 1.0.0 py27_0 <unknown>
xz 5.2.4 h1de35cc_4
yaml 0.1.6 0
zeromq 4.1.3 0
zict 0.1.3 py27h5fff8b1_0
zlib 1.2.11 0 conda-forge
pip list
Package Version
---------------------------------- ------------
abstract-rendering 0.5.1
ads 0.12.3
alabaster 0.7.6
anaconda-client 1.5.1
anaconda-navigator 1.3.1
APLpy 1.1.1
appnope 0.1.0
appscript 1.0.1
argcomplete 0.8.9
asdf 1.2.1
astro-gala 0.1.3
astroid 1.4.7
astroML 0.3
astroplan 0.2
astropy 2.0.3
astropy-helpers 1.1.1
astroquery 0.3.7
astroscrappy 1.0.5
Babel 2.1.1
backports-abc 0.4
backports.functools-lru-cache 1.5
backports.shutil-get-terminal-size 1.0.0
backports.ssl-match-hostname 3.4.0.2
basemap 1.1.0
bcolz 0.9.0
beautifulsoup4 4.5.1
binstar 0.11.0
bitarray 0.8.1
blaze 0.8.0
bleach 1.5.0
blinker 1.4
blz 0.6.2
bokeh 0.12.16
boto 2.38.0
Bottleneck 1.0.0
brewer2mpl 1.4.1
Cartopy 0.16.0
ccdproc 1.2.0
cdecimal 2.3
certifi 2018.4.16
cffi 1.9.1
chardet 3.0.4
chest 0.2.3
click 6.6
click-plugins 1.0.3
cligj 0.4.0
cloudpickle 0.2.2
cluster-lensing 0.1.2
clyent 0.3.4
colorama 0.3.3
conda 4.5.4
conda-build 1.14.1
configobj 5.0.6
configparser 3.5.0b2
corner 2.0.1
coverage 3.7.1
cryptography 1.7.1
cycler 0.10.0
Cython 0.24
cytoolz 0.7.3
d2to1 0.2.12.post1
dask 0.16.0
datashape 0.4.5
decorator 4.0.10
dill 0.2.7.1
distributed 1.20.2
docutils 0.12
emcee 2.2.1
enum34 1.1.6
everest-pipeline 2.0.8
extinction 0.3.0
fastcache 1.0.2
feedgenerator 1.7
Flask 0.10.1
funcsigs 1.0.2
functools32 3.2.3.post2
future 0.16.0
futures 3.2.0
galpy 1.2
gammapy 0.4
gatspy 0.3
george 0.2.1
gevent 1.0.1
gevent-websocket 0.9.3
ginga 2.6.1
glue-core 0.12.4
glue-vispy-viewers 0.9.2
gPhoton 1.28.2
greenlet 0.4.7
grin 1.2.1
gwcs 0.7
h5py 2.7.0
halotools 0.4
healpy 1.8.6
HeapDict 1.0.0
html5lib 0.9999999
httpretty 0.8.14
idna 2.6
imageio 2.2.0
imagesize 0.7.1
imexam 0.7.0
ipaddress 1.0.7
ipykernel 4.6.1
ipython 5.3.0
ipython-genutils 0.1.0
ipywidgets 6.0.0
itsdangerous 0.24
jdcal 1.0
jedi 0.9.0
Jinja2 2.8
jsonschema 2.4.0
jupyter-client 4.2.2
jupyter-core 4.1.0
k2flix 1.1.dev0
K2fov 6.2.0
k2plr 0.2.5
keyring 9.0
kiwisolver 1.0.1
lazy-object-proxy 1.2.1
llvmlite 0.9.0
lmfit 0.9.5
locket 0.2.0
lxml 4.1.1
maltpynt 2.0
Markdown 2.6.9
MarkupSafe 0.23
matplotlib 2.2.2
mistune 0.7.2
mock 2.0.0
modernize 0.5
montage-wrapper 0.9.8
mpld3 0.2
mpldatacursor 0.6.2
mpmath 0.19
msgpack-python 0.4.8
multipledispatch 0.4.7
munch 2.0.4
naima 0.8
nbconvert 4.2.0
nbformat 4.0.1
networkx 1.11
nltk 3.0.3
nose 1.3.7
notebook 5.0.0
numba 0.24.0
numexpr 2.6.4
numpy 1.14.0
numpydoc 0.6.0
odo 0.3.2
olefile 0.45.1
omnifit 0.2.1
openpyxl 1.8.5
OWSLib 0.16.0
packaging 17.1
palpy 1.6.0
pandas 0.21.1
pandas-qt 0.1.2
partd 0.3.8
path.py 0.0.0
pathlib2 2.2.1
patsy 0.4.1
pbr 3.1.1
pelican 3.6.3
pep8 1.6.2
pexpect 4.0.1
phoebe 2.0b0
photutils 0.3
pickleshare 0.5
Pillow 4.2.1
pip 10.0.1
plotly 2.0.15
ply 3.6
probableparsing 0.0.1
prompt-toolkit 1.0.14
protobuf 3.5.1
psutil 3.3.0
ptyprocess 0.5
py 1.5.3
pyasn1 0.1.9
PyAudio 0.2.7
pycodestyle 2.3.1
pycosat 0.6.3
pycparser 2.14
pycrypto 2.6.1
pycurl 7.19.5.1
pydl 0.5.3
pyephem 3.7.5.3
pyepsg 0.3.2
pyfits 3.4
pyflakes 1.0.0
Pygments 2.1.3
pyketools 3.0b3
pylint 1.5.4
pymc 2.3.6
PyOpenGL 3.1.1a1
pyOpenSSL 16.2.0
pyparsing 2.2.0
PyPDF2 1.26.0
pyproj 1.9.5.1
pyqtgraph 0.9.10
pyregion 2.0
pyshp 1.2.12
PySocks 1.6.8
pysqlite 2.8.3
pysyzygy 0.0.1
pytest 2.9.1
pytest-cov 2.1.0
pytest-qt 1.2.2
python-cpl 0.7.2
python-crfsuite 0.9.2
python-dateutil 2.6.0
pytz 2017.2
pyvo 0.5.1
PyWavelets 0.5.2
PyYAML 3.11
pyzipcode 1.0
pyzmq 15.2.0
QtAwesome 0.4.1
qtconsole 4.3.0
QtPy 1.2.0
redis 2.10.3
reproject 0.3.2
requests 2.11.0
rope 0.9.4
ruamel-yaml -VERSION
runipy 0.1.3
scandir 1.5
scikit-image 0.13.0
scikit-learn 0.19.1
scipy 1.0.0
seaborn 0.7.1
setuptools 38.4.0
setuptools-git 1.1
Shapely 1.6.3
simplegeneric 0.8.1
singledispatch 3.4.0.3
six 1.10.0
sncosmo 1.4.0
snowballstemmer 1.2.0
sockjs-tornado 1.0.1
sortedcontainers 1.5.7
spectral-cube 0.4.0
specutils 0.2.2
spherical-geometry 1.0.6
Sphinx 1.4.6
sphinx-rtd-theme 0.1.7
spyder 3.2.8
SQLAlchemy 1.0.12
statsmodels 0.8.0
stsci.distutils 0.3.7
subprocess32 3.2.7
sympy 1.0
tables 3.2.0
tblib 1.3.2
tensorflow 1.4.0
tensorflow-tensorboard 0.4.0rc3
terminado 0.5
titlecase 0.12.0
toolz 0.8.2
tornado 4.5.2
tqdm 4.14.0
traitlets 4.3.2
ujson 1.33
unicodecsv 0.9.4
Unidecode 0.4.19
urllib3 1.22
usaddress 0.5.10
wcsaxes 0.9
wcwidth 0.1.7
webencodings 0.5
Werkzeug 0.12.2
wheel 0.29.0
widgetsnbextension 2.0.0
wikipedia 1.4.0
wrapt 1.10.8
xlrd 1.1.0
XlsxWriter 0.7.3
xlwings 0.3.5
xlwt 1.0.0
zict 0.1.3
If you change your longitudes to be -pi -> pi rather than 0 -> 2*pi, contourf works as expected. Weird that it would work differently in the two different methods depending on the longitudes.
This is the change I made:
lon = np.linspace(0,2*np.pi,200) - np.pi
The default central longitude for PlateCarree is 0, so I guess it makes sense that the input range should be -pi <-> pi.
Thanks for your input!
Subtracting pi did work for that particular example, but it does not fix the problem generally for other data. For instance, if I change to
lon = np.linspace(0,2*np.pi,200)-np.pi
and add
drm = -1.*drm
before plotting, I now get:

That is curious indeed! It looks like the issues arise when the drm values are close to zero. Following a suggestion from this basemap issue
drm[np.abs(drm) < 1.e-6] = 0.
Setting small values of drm to zero directly worked for me in all cases. Even with lons 0 -> 2*pi. So, my original suggestion had nothing to do with the problem it looks like. I'm still not sure why this is happening, but hope this helps you out anyways.
Yes, that does seem to work in most cases, but I have to tune that "close to zero" limit depending on the situation.
I think I actually found a simple fix: setting the contour line values explicitly instead of just specifying the number.
I have no idea why this makes a difference. Inspecting the levels property returned in the contourf object, nothing looks odd. Things work for me when I specify those exact levels explicitly.
I'm experiencing the same on some geo data plots. In some cases (apparently casually but reproducible and linked with data resolution) the negative contourf disappear from the plots. Opening the produced pdf files with Inkscape, I found that the negative contourf are in effect produced but then hided by other levels. I'm attaching the original plot and the reconstructed one, in which I just changed the order of levels inside Inkscape. It appears to be a problem with the zorder of contourf patches..
ps. I tried outputting different filetypes, but the problem persists, and for the same datasets bug.pdf .
It's not really a zorder problem; some paths just get transformed to fill the whole area when they shouldn't.
So, I can reproduce this problem if I pass levels = np.arange(-42, 43, 2) / 100, but not levels = np.arange(-0.42, 0.43, 0.02). The difference being that, due to floating point addition, in the latter case the '0' contour level is actually 3.88578059e-16. So the path is just ever so slightly different enough to not trigger the bad transform.
The '0' contour level path is generally one that fills the entire Plate Carree space (see below), with some holes cut out of it for the other contours. Because of that, I think the problem is fairly similar to #1149 and #146 as it has to do with polygons that fill the entire space and need to be stitched back together, but aren't correctly.

Well, I'm a bit confused now, because I can extract the path used to plot the 0-contour, and plot it on a new figure, and it appears fine:
cf = ax0.contourf(lon*180/np.pi, lat*180/np.pi, drm, levels,
transform=ccrs.PlateCarree(), cmap='seismic')
for i, pc in enumerate(cf.collections):
if i == 20:
path = pc.get_paths()[0]
fig1 = plt.figure()
ax1 = fig1.add_subplot(1, 1, 1, projection=ccrs.Orthographic(0, 30))
ax1.add_patch(
matplotlib.patches.PathPatch(path, transform=ccrs.PlateCarree()))
ax1.set_global()

which naturally makes it really hard to debug.
@QuLogic I think you must have had your 'levels' fix in there from before. When I use your code I get what you were saying earlier with the zero level taking up the entire area. The reason the high regions still show up is because of the drawing order. I agree with all of your assessments of related issues and clipping/patching the paths together being the issue. I don't have a solution for that though.
Here is a contained example to reproduce the issues for copy.paste purposes.
import matplotlib
import matplotlib.pyplot as plt
import numpy as np
import scipy.special as sp
import cartopy
import cartopy.crs as ccrs
# Compute spherical harmonic pattern
l=4; m=3
lon = np.linspace(0,2*np.pi,200)
lat = np.linspace(-np.pi/2,np.pi/2,500)
colat = lat+np.pi/2
d = np.zeros((len(lon),len(colat)),dtype = np.complex64)
meshed_grid = np.meshgrid(lon, lat)
lat_grid = meshed_grid[1]
lon_grid = meshed_grid[0]
for j, yy in enumerate(colat):
for i, xx in enumerate(lon):
d[i,j] = sp.sph_harm(m,l,xx,yy)
drm = np.transpose(np.real(d))
#Plot example with contour() and contourf()
fig, (ax1, ax2) = plt.subplots(1, 2, subplot_kw={'projection': ccrs.Orthographic(0, 30)})
levels = np.arange(-42, 43, 2) / 100
#levels = np.arange(-0.42, 0.43, 0.02)
#contour() works as expected
ax1.contour(lon*180/np.pi,lat*180/np.pi,drm,levels,
transform=ccrs.PlateCarree(),cmap='seismic')
ax1.relim()
ax1.autoscale_view()
#contourf() doesn't
cf = ax2.contourf(lon*180/np.pi,lat*180/np.pi,drm,levels,
transform=ccrs.PlateCarree(),cmap='seismic')
fig1 = plt.figure()
ax1 = fig1.add_subplot(1, 2, 1, projection=ccrs.Orthographic(0, 30))
ax2 = fig1.add_subplot(1, 2, 2, projection=ccrs.PlateCarree())
for i, pc in enumerate(cf.collections):
if i == 20:
path = pc.get_paths()[0]
ax1.add_patch(
matplotlib.patches.PathPatch(path, transform=ccrs.PlateCarree()))
ax1.set_global()
ax2.add_patch(
matplotlib.patches.PathPatch(path, transform=ccrs.PlateCarree()))
ax2.set_global()
@greglucas Ah right, sorry I forgot, you must copy it to make it fix itself: path = pc.get_paths()[0].deepcopy(), which is essentially the same as saving to a file and loading again.
Hi, does someone have any news on this? I moved to the Stereographic projection to avoid the bug, but would be great to go back to the Orthographic..
I ended up using pcolormesh for my application instead of contourf to avoid this issue. Not exactly the same, but at least I can get an Orthographic projection this way.
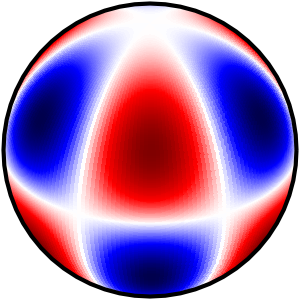
Hi,
a short update on this. I initially thought the Stereographic would work, but actually I find the same bug (i.e. negative contourf not drawn) for the Stereographic and the Nearside projections. The bug appears very often (more than 50% of the cases) when a contourf is crossing the 0 value, which makes that contourf spread to cover the full image. It does not appear when 0 is one of the levels.
I think this is a very serious bug. This makes it impossible to reliably produce filled contours when looking at the poles. Is there a timeline for solving this? It has been more than one year now..
@fedef17 If you can reduce down to a self-contained example that reproduces the problem, that would be helpful. We have one of those above, but it would be good to collect several, since I'm sure we're dealing with edge cases in the current algorithm.
It seems that the issue described in the first example can be managed by the workaround presented in issue #1421 as this seems to be a case of overlapping data.
The results obtained with the code below:

import matplotlib.pyplot as plt
import numpy as np
import numpy.ma as ma
import scipy.special as sp
import cartopy.crs as ccrs
def z_masked_overlap(axe, X, Y, Z, source_projection=None):
"""
for data in projection axe.projection
find and mask the overlaps (more 1/2 the axe.projection range)
X, Y either the coordinates in axe.projection or longitudes latitudes
Z the data
operation one of 'pcorlor', 'pcolormesh', 'countour', 'countourf'
if source_projection is a geodetic CRS data is in geodetic coordinates
and should first be projected in axe.projection
X, Y are 2D same dimension as Z for contour and contourf
same dimension as Z or with an extra row and column for pcolor
and pcolormesh
return ptx, pty, Z
"""
if not hasattr(axe, 'projection'):
return X, Y, Z
if not isinstance(axe.projection, ccrs.Projection):
return X, Y, Z
if len(X.shape) != 2 or len(Y.shape) != 2:
return X, Y, Z
if (source_projection is not None and
isinstance(source_projection, ccrs.Geodetic)):
transformed_pts = axe.projection.transform_points(
source_projection, X, Y)
ptx, pty = transformed_pts[..., 0], transformed_pts[..., 1]
else:
ptx, pty = X, Y
with np.errstate(invalid='ignore'):
# diagonals have one less row and one less columns
diagonal0_lengths = np.hypot(
ptx[1:, 1:] - ptx[:-1, :-1],
pty[1:, 1:] - pty[:-1, :-1]
)
diagonal1_lengths = np.hypot(
ptx[1:, :-1] - ptx[:-1, 1:],
pty[1:, :-1] - pty[:-1, 1:]
)
to_mask = (
(diagonal0_lengths > (
abs(axe.projection.x_limits[1]
- axe.projection.x_limits[0])) / 2) |
np.isnan(diagonal0_lengths) |
(diagonal1_lengths > (
abs(axe.projection.x_limits[1]
- axe.projection.x_limits[0])) / 2) |
np.isnan(diagonal1_lengths)
)
# TODO check if we need to do something about surrounding vertices
# add one extra colum and row for contour and contourf
if (to_mask.shape[0] == Z.shape[0] - 1 and
to_mask.shape[1] == Z.shape[1] - 1):
to_mask_extended = np.zeros(Z.shape, dtype=bool)
to_mask_extended[:-1, :-1] = to_mask
to_mask_extended[-1, :] = to_mask_extended[-2, :]
to_mask_extended[:, -1] = to_mask_extended[:, -2]
to_mask = to_mask_extended
if np.any(to_mask):
Z_mask = getattr(Z, 'mask', None)
to_mask = to_mask if Z_mask is None else to_mask | Z_mask
Z = ma.masked_where(to_mask, Z)
return ptx, pty, Z
# Compute spherical harmonic pattern
l=4; m=3
lon = np.linspace(0,2*np.pi,200)
lat = np.linspace(-np.pi/2,np.pi/2,500)
colat = lat+np.pi/2
d = np.zeros((len(lon),len(colat)),dtype = np.complex64)
meshed_grid = np.meshgrid(lon, lat)
lat_grid = meshed_grid[1]
lon_grid = meshed_grid[0]
for j, yy in enumerate(colat):
for i, xx in enumerate(lon):
d[i,j] = sp.sph_harm(m,l,xx,yy)
drm = np.transpose(np.real(d))
#Plot example with contour() and contourf()
fig, ((ax1, ax2), (ax3, ax4)) = plt.subplots(2, 2,
subplot_kw={'projection': ccrs.Orthographic(0, 30)})
#contour() works as expected
ax1.contour(lon*180/np.pi,lat*180/np.pi,drm,51,
transform=ccrs.PlateCarree(),cmap='seismic')
ax1.relim()
ax1.autoscale_view()
#contourf() doesn't
ax2.contourf(lon*180/np.pi,lat*180/np.pi,drm,51,
transform=ccrs.PlateCarree(),cmap='seismic')
ax2.relim()
ax2.autoscale_view()
lons, lats = np.meshgrid(lon*180/np.pi, lat*180/np.pi)
# mask the overlaps
X, Y, masked_drm = z_masked_overlap(
ax4, lons, lats, drm,
source_projection=ccrs.Geodetic())
ax3.contour(X ,Y ,masked_drm, 51,
cmap='seismic')
ax3.relim()
ax3.autoscale_view()
ax3.set_title('after z_masked_overlap workaround')
ax4.contourf(X ,Y ,masked_drm, 51,
cmap='seismic')
ax4.relim()
ax4.autoscale_view()
ax4.set_title('after z_masked_overlap workaround')
plt.tight_layout()
plt.savefig('img/sphericalcontours.png')
@fedef17 If you can reduce down to a self-contained example that reproduces the problem, that would be helpful. We have one of those above, but it would be good to collect several, since I'm sure we're dealing with edge cases in the current algorithm.
Hi @dopplershift, here is a self-contained example from my work to reproduce the problem. I agree with previous comments that this is rather serious from a user's perspective, and it would be good to fix it soon. I don't have the expertise to debug cartopy but I'm happy to help where I can.
For me the problem only occurs under the following conditions:
- Using projection cartopy.crs.NorthPolarStereo (not SouthPolarStereo)
- Contour levels are sufficiently large
- Data has been prepared using cartopy.util.add_cyclic_point
I have tested with cartopy version 0.18.0b1.
Below the plot demonstrating the problem together with the code that produces the plot. Data file (~200kB, NetCDF4) is attached.

import matplotlib.pyplot as plt
import xarray as xr
import cartopy
import cartopy.crs as ccrs
from cartopy.util import add_cyclic_point
# load data
fig = plt.figure(figsize=(12, 4))
da = xr.open_dataarray('testdata.nc')
lats = da.latitude.values
lons_nocyclic = da.longitude.values
data_nocyclic = da.values
data, lons = add_cyclic_point(da.values, coord=lons_nocyclic)
# negative contours not shown for these contour levels
nok_levels = [-0.4, -0.2, 0.2, 0.4]
# negative contours are correctly shown for these contour levels
ok_levels = [-.4, -.3, -0.2, -0.1, 0.1, 0.2, .3, .4]
# keyword arguments for contourf and colorbar
conargs = dict(transform=ccrs.PlateCarree(), cmap='RdBu_r', extend='both')
cbargs = dict(orientation='horizontal', pad=0.05, extend='both')
# 1) problem: negative contours not shown
plt.subplot(1, 3, 1, projection=ccrs.NorthPolarStereo())
c1 = plt.contourf(lons, lats, data, nok_levels, **conargs)
plt.colorbar(c1, **cbargs)
plt.title('(a) missing negative contours')
# 2) problem disappears with slightly different contour levels
plt.subplot(1, 3, 2, projection=ccrs.NorthPolarStereo())
c2 = plt.contourf(lons, lats, data, ok_levels, **conargs)
plt.colorbar(c2, **cbargs)
plt.title('(b) change levels: no problem')
# 3) problem disappears with slightly different contour levels
plt.subplot(1, 3, 3, projection=ccrs.NorthPolarStereo())
c3 = plt.contourf(lons_nocyclic, lats, data_nocyclic, nok_levels, **conargs)
plt.colorbar(c3, **cbargs)
plt.title('(c) no cyclic point: no problem')
# add decorations to plots and save
for ax in plt.gcf().axes:
if type(ax) == cartopy.mpl.geoaxes.GeoAxesSubplot:
ax.set_extent((-180.0, 180.0, 45.0, 90.0), ccrs.PlateCarree())
ax.gridlines()
ax.coastlines()
fig.savefig('cartopy_test.png')
I think that last problem should be fixed by @htonchia's idea here, though I'm not sure it'll fix the original problem.
I also came across this problem recently. Here's another example that reproduces it:
import numpy as np
import matplotlib.pyplot as plt
import cartopy.crs as ccrs
N = 10
state = np.random.RandomState(23)
fig = plt.figure(figsize=(5, 2.5))
ax = fig.add_subplot(projection=ccrs.Mollweide())
ax.coastlines()
x = np.linspace(-180, 180, N)
y = np.linspace(-90, 90, N)
z = state.rand(N, N) * 10 - 5
m = ax.contourf(
x, y, z,
transform=ccrs.PlateCarree(),
cmap='RdBu_r',
vmin=-5,
vmax=5,
)
fig.colorbar(m, ax=ax)
And the resulting output:

It seems to depend on the figure size, dataset, and projection -- sometimes the problem appears, sometimes it doesn't. I had to randomly try different np.random.RandomState seeds to generate this result. Very very strange.
I've just come across this too, in cartopys 0.17, 0.18, 0.20.1.
My code: (the data file temp.nc is in the attached temp.zip)
import numpy as np
import cartopy
import cartopy.crs as ccrs
import iris
import iris.quickplot as qplt
import matplotlib.pyplot as plt
mycube = iris.load_cube("temp.nc")
proj = ccrs.Orthographic(central_longitude=0.0, central_latitude=90.0)
# This works correctly (Figure 1):
qplt.contourf(mycube,np.linspace(-9,0,11),axes=plt.axes(projection=proj)) ; qplt.plt.gca().coastlines() ; qplt.show()
# This plots incorrectly (Figure 2):
qplt.contourf(mycube,np.linspace(-9,0,10),axes=plt.axes(projection=proj)) ; qplt.plt.gca().coastlines() ; qplt.show()
# There is no problem on the Plate Carree projection:
qplt.contourf(mycube,np.linspace(-9,0,11)) ; qplt.plt.gca().coastlines() ; qplt.show()
qplt.contourf(mycube,np.linspace(-9,0,10)) ; qplt.plt.gca().coastlines() ; qplt.show()
# Also works without iplt: (thanks @rcomer !)
axes = plt.axes(projection=proj)
plt.contourf(
mycube.coord("longitude").points,
mycube.coord("latitude").points,
mycube.data,
levels=np.linspace(-9, 0, 10),
transform=ccrs.PlateCarree(),
)
axes.coastlines()
plt.colorbar(orientation="horizontal")
plt.show()
Plots:


If I make Figure 2 as a svg and load it into Inkscape, I can confirm that the other contour levels are present, just underneath the zero-layer.
In my case, setting the cube data values near zero to be exactly zero doesn't help;
and np.linspace(-9,0,11) and np.linspace(-9,0,10) both seem to have zeros at exactly zero (rather than somethingE-16)
I am currently having this issue in Cartopy 0.21.1. Here is my code:
import numpy as np
import math
import csv
import matplotlib.pyplot as plt
import cartopy.crs as ccrs
import cartopy.feature as cf
resY = 360
resX = 720
extent_globe = np.radians([0,360,-90,90])
grid_lon = np.linspace(extent_globe[0], extent_globe[1], resX)
grid_lat = np.linspace(extent_globe[2], extent_globe[3], resY)
grid_lons, grid_lats = np.meshgrid(grid_lon, grid_lat)
part1 = np.square(np.tan(math.pi / 3)) / np.square(np.tan(grid_lats))
btheta = np.where(grid_lats > 0, part1 * np.exp(1 - part1), 0)
cpart = 0.6 * 2 * math.pi * np.cos(1 * grid_lons)
forcing = btheta * cpart
fig = plt.figure()
ax1 = plt.subplot(projection=ccrs.Orthographic(central_latitude=90))
cs = ax1.contourf(np.rad2deg(grid_lons), np.rad2deg(grid_lats), forcing, cmap='RdBu_r', transform=ccrs.PlateCarree(), levels=np.mgrid[-10:10:21j])
plt.title(f'Wavenumber 1 Forcing')
cbar = fig.colorbar(cs)
plt.show()
In this case, it shows the negative contours if I replace forcing with -forcing, but if I replace np.cos(1 * grid_lons) with np.cos(2 * grid_lons) then the negative sign does not reveal the negative contours. The previously suggested solutions of changing the longitude range to be 0 to 360 and zeroing out small values did not fix the error.