Contour plot breaks with certain contour levels
Obscure error when running ACME Diags on output from a recent simulation:
/export/golaz1/ACME_diags/20170926.FCT2.A_WCYCL1850S.ne30_oECv3.anvil
Variable: LWCF
Selected region: global
no domain selector
No handlers could be found for logger "shapely.geos"
Traceback (most recent call last):
File "/export/golaz1/conda/envs/acme_diags_env/bin/acme_diags_driver.py", line 4, in <module>
__import__('pkg_resources').run_script('acme-diags==1.0.0', 'acme_diags_driver.py')
File "/export/golaz1/conda/envs/acme_diags_env/lib/python2.7/site-packages/pkg_resources/__init__.py", line 742, in run_script
self.require(requires)[0].run_script(script_name, ns)
File "/export/golaz1/conda/envs/acme_diags_env/lib/python2.7/site-packages/pkg_resources/__init__.py", line 1503, in run_script
exec(code, namespace, namespace)
File "/export/golaz1/conda/envs/acme_diags_env/lib/python2.7/site-packages/acme_diags-1.0.0-py2.7.egg-info/scripts/acme_diags_driver.py", line 122, in <module>
parameters = cdp.cdp_run.serial(run_diag, parameters)
File "/export/golaz1/conda/envs/acme_diags_env/lib/python2.7/site-packages/cdp/cdp_run.py", line 12, in serial
results.append(func(p))
File "/export/golaz1/conda/envs/acme_diags_env/lib/python2.7/site-packages/acme_diags-1.0.0-py2.7.egg-info/scripts/acme_diags_driver.py", line 59, in run_diag
single_result = module.run_diag(parameters)
File "/export/golaz1/conda/envs/acme_diags_env/lib/python2.7/site-packages/acme_diags/driver/lat_lon_driver.py", line 224, in run_diag
mv1_domain, diff, metrics_dict, parameter)
File "/export/golaz1/conda/envs/acme_diags_env/lib/python2.7/site-packages/acme_diags/plot/__init__.py", line 43, in plot
plot_fcn(ref, test, diff, metrics_dict, parameter)
File "/export/golaz1/conda/envs/acme_diags_env/lib/python2.7/site-packages/acme_diags/plot/cartopy/lat_lon_plot.py", line 149, in plot
(None, parameter.diff_title, None), parameter, stats=(max3, mean3, min3, r, c))
File "/export/golaz1/conda/envs/acme_diags_env/lib/python2.7/site-packages/acme_diags/plot/cartopy/lat_lon_plot.py", line 61, in plot_panel
extend='both',
File "/export/golaz1/conda/envs/acme_diags_env/lib/python2.7/site-packages/cartopy/mpl/geoaxes.py", line 1339, in contourf
if col.get_paths()])
File "/export/golaz1/conda/envs/acme_diags_env/lib/python2.7/site-packages/matplotlib/collections.py", line 214, in get_datalim
paths = [transform.transform_path_non_affine(p) for p in paths]
File "/export/golaz1/conda/envs/acme_diags_env/lib/python2.7/site-packages/matplotlib/transforms.py", line 2386, in transform_path_non_affine
return self._a.transform_path_non_affine(path)
File "/export/golaz1/conda/envs/acme_diags_env/lib/python2.7/site-packages/cartopy/mpl/geoaxes.py", line 187, in transform_path_non_affine
geom, self.source_projection)
File "/export/golaz1/conda/envs/acme_diags_env/lib/python2.7/site-packages/cartopy/crs.py", line 175, in project_geometry
return getattr(self, method_name)(geometry, src_crs)
File "/export/golaz1/conda/envs/acme_diags_env/lib/python2.7/site-packages/cartopy/crs.py", line 330, in _project_polygon
return self._rings_to_multi_polygon(rings, is_ccw)
File "/export/golaz1/conda/envs/acme_diags_env/lib/python2.7/site-packages/cartopy/crs.py", line 541, in _rings_to_multi_polygon
if prep_polygon.contains(interior_ring):
File "/export/golaz1/conda/envs/acme_diags_env/lib/python2.7/site-packages/shapely/impl.py", line 37, in wrapper
return func(*args, **kwargs)
File "/export/golaz1/conda/envs/acme_diags_env/lib/python2.7/site-packages/shapely/prepared.py", line 45, in contains
return bool(self.impl['prepared_contains'](self, other))
File "/export/golaz1/conda/envs/acme_diags_env/lib/python2.7/site-packages/shapely/predicates.py", line 18, in __call__
self._check_topology(err, this, other)
File "/export/golaz1/conda/envs/acme_diags_env/lib/python2.7/site-packages/shapely/topology.py", line 34, in _check_topology
if not geom.is_valid:
AttributeError: 'PreparedGeometry' object has no attribute 'is_valid'
I isolated the problem. It seems to be related to plotting MAM against CERES EBAF TOA 4.0. Plotting MAM against 2.8 works. Could there be something wrong with CERES EBAF TOA 4.0 MAM file?
Here is my JSON file:
{
"set5": [
{
"sets": ["lat_lon"],
"case_id": "CERES-EBAF-TOA-v4.0",
"variables": ["LWCF"],
"ref_name": "ceres_ebaf_toa_v4.0",
"reference_name": "CERES-EBAF Jan 2000-Dec 2015",
"seasons": ["MAM"],
"contour_levels": [0, 10, 20, 30, 40, 50, 60, 70, 80],
"diff_levels": [-35, -30, -25, -20, -15, -10, -5, -2, 2, 5, 10, 15, 20, 25, 30, 35]
}
]
}
@golaz Since this seems like a problem related to matplotlib, did you want to try just running the aforementioned diags using vcs? It seems weird that the plotting is causing the errors.
Hi @golaz where is the data? on which machine? Seems like it's not on Nersc. I'm hoping to look at the data. Seems like, below line is causing the problem.
File "/export/golaz1/conda/envs/acme_diags_env/lib/python2.7/site-packages/acme_diags/plot/cartopy/lat_lon_plot.py", line 149, in plot (None, parameter.diff_title, None), parameter, stats=(max3, mean3, min3, r, c))
Maybe it's the metrics ...I'm hoping to reproduce the problem.
@zshaheen: I tried with VCS and it works
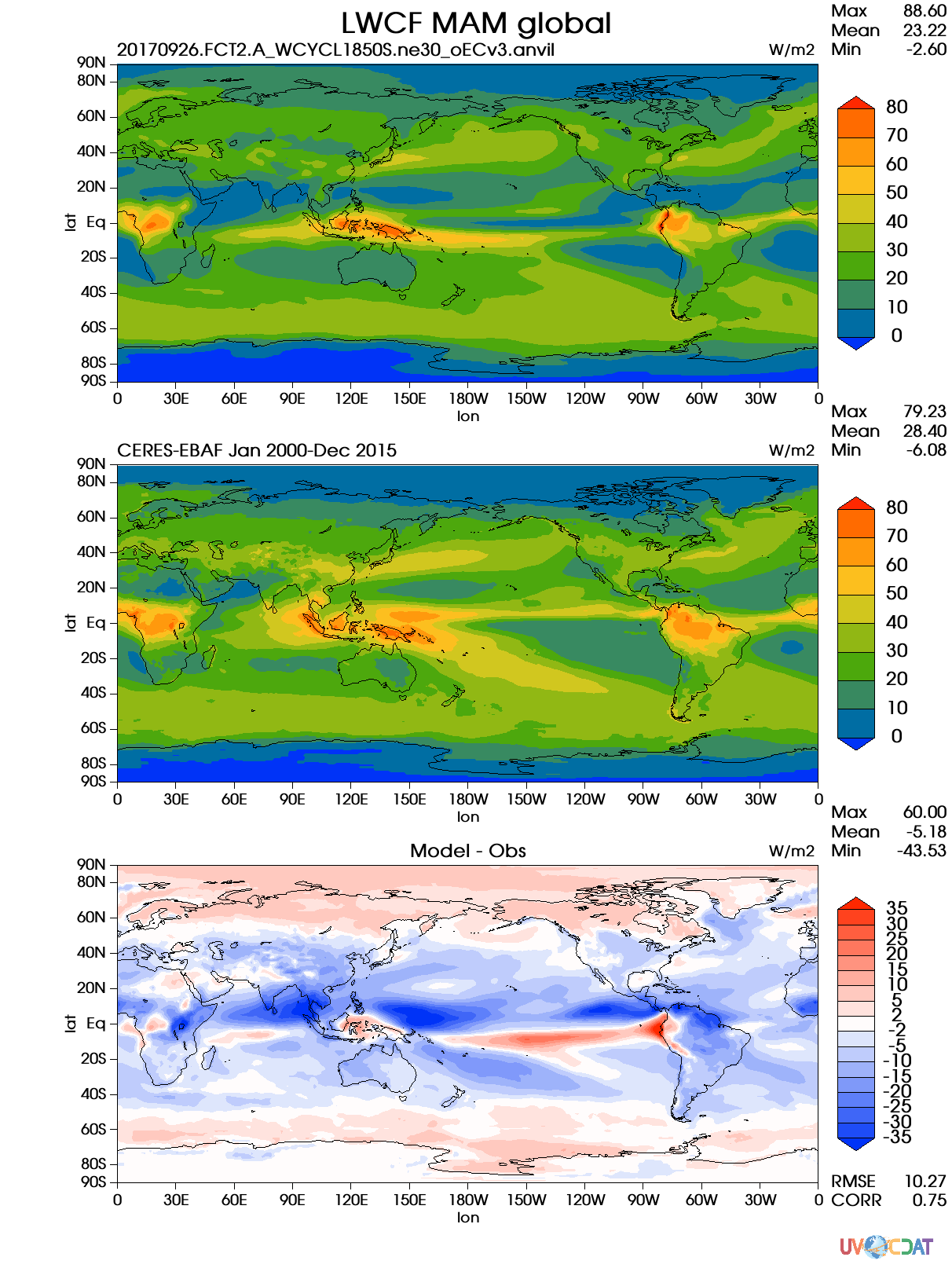
@golaz @chengzhuzhang So it seems like the metrics aren't an issue, because they are just floating point numbers, nothing weird.
@chengzhuzhang : I'm seeing this error on acme1.
Model data:
/export/golaz1/ACME_diags/20170926.FCT2.A_WCYCL1850S.ne30_oECv3.anvil/climo/0001-0030
Obs:
/p/cscratch/acme/data/obs_for_acme_diags
@golaz
Thank you for helping us ruling out the metrics problem. I will try to reproduce this error and trouble shooting it.
I was trying to do additional debugging by creating a development environment, but then ran into another issue: https://github.com/ACME-Climate/acme_diags/issues/99
The problem goes away when slightly tweaking the specified contour levels for the difference plot. For example, the following works:
{
"set5": [
{
"sets": ["lat_lon"],
"case_id": "CERES-EBAF-TOA-v4.0",
"variables": ["LWCF"],
"ref_name": "ceres_ebaf_toa_v4.0",
"reference_name": "CERES-EBAF Jan 2000-Dec 2015",
"seasons": ["MAM"],
"contour_levels": [0, 10, 20, 30, 40, 50, 60, 70, 80],
"diff_levels": [-35, -25, -15, -5, -2, 2, 5, 15, 25, 35]
}
]
}
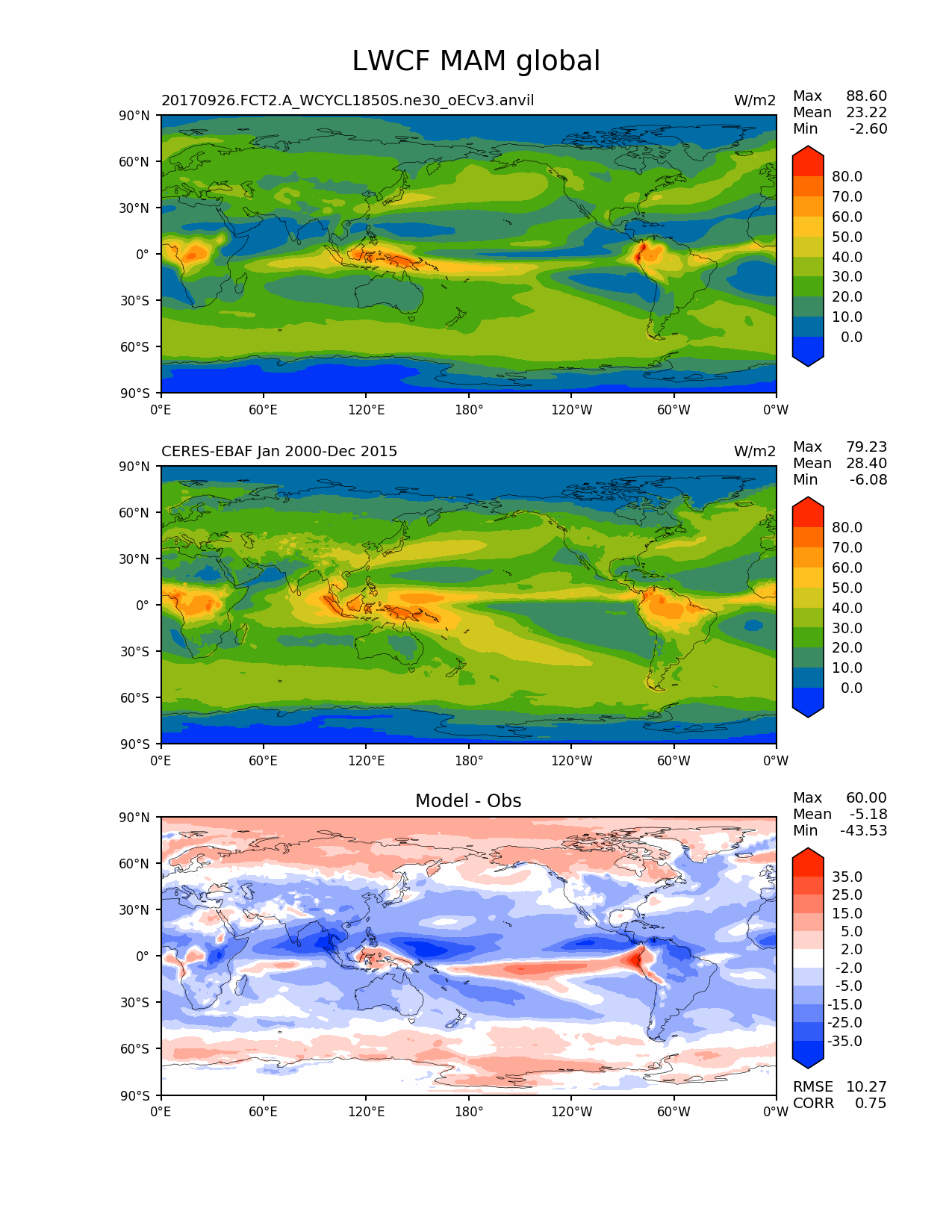
My current best hypothesis is that this is a cartopy bug. The behavior is very similar to an older cartopy bug (https://github.com/SciTools/cartopy/issues/811). It supposedly was corrected in version 0.15.1 that we are using, but maybe not quite (?)
Maybe we need a simple test illustrating the problem.
@golaz Hi Chris,
Would you give me permission for: /export/golaz1/ACME_diags/20170926.FCT2.A_WCYCL1850S.ne30_oECv3.anvil/climo/0001-0030/
Thank you!
@golaz Zeshawn helped to moved the data, no permission needed now..
I can totally reproduce this issue. A cartopy bug needs to be fixed...
@chengzhuzhang We should isolate this bug from our software and write a short script that demonstrates this. Then we can show this to the maintainers of cartopy.
@zshaheen I agree with you. It seems to me a random case, since it only happens to one variable (derived) for one season so far. It's wired. I'm thinking in our software, we should raise this error but continue with some remedy. definitely not stop the run.
@chengzhuzhang and @zshaheen: I like your two prongs approach:
- Implement a work-around so that ACME diags continues even if a single plot fails (we also need something like this for https://github.com/ACME-Climate/acme_diags/issues/95).
- If possible, create a simple test case that isolates the problem and submit it as a bug report to cartopy. I wonder if we can make use of the option to save the regridded netCDF files and then create a simple one-panel plot that would trigger the bug.
@golaz Yes! I actually also thought save_neccdf should be used right here. I'm creating the simple sample.
@chengzhuzhang and @zshaheen: I encountered a similar problem on NERSC yesterday with a different set of climatology files:
/global/cscratch1/sd/golaz/ACME_simulations/20171011.beta2_FCT2-rt02_branch.A_WCYCL1850S.ne30_oECv3_ICG.edison/pp/clim_rgr/0051-0060
This time, the error occurred when creating SWCF MAM figures with EBAF 2.8. The version of ACME diags I was using included PR https://github.com/ACME-Climate/acme_diags/pull/103, so additional changes are probably needed to recover from this specific type of failure.
@golaz Jill and I came up with a solution that should step around most of these issues. We'll present this to you tomorrow. Since this is an issue with a library we use, stepping around this probably the only thing we can do.
@golaz With this new error Chris, did the diagnostics continue past the error, or did it prematurely break?
Also, can you post the stack trace if possible?
It broke prematurely. Traceback looks very similar to the one above; should be reproducible with the climo files mentioned above.
test file: /global/cscratch1/sd/golaz/ACME_simulations/20171011.beta2_FCT2-rt02_branch.A_WCYCL1850S.ne30_oECv3_ICG.edison/pp/clim_rgr/0051-0060/20171011.beta2_FCT2-rt02_branch.A_WCYCL1850S.ne30_oECv3_ICG.edison_MAM_005103_006005_climo.nc
reference file: /global/project/projectdirs/acme/acme_diags/obs_for_acme_diags/ceres_ebaf_toa_v2.8_MAM_200103_201605_climo.nc
Variable: SWCF
Selected region: global
no domain selector
No handlers could be found for logger "shapely.geos"
Traceback (most recent call last):
File "/global/project/projectdirs/acme/golaz/conda/envs/acme_diags_dev/bin/acme_diags_driver.py", line 4, in <module>
__import__('pkg_resources').run_script('acme-diags==1.0.0', 'acme_diags_driver.py')
File "/global/project/projectdirs/acme/golaz/conda/envs/acme_diags_dev/lib/python2.7/site-packages/pkg_resources/__init__.py", line 748, in run_script
self.require(requires)[0].run_script(script_name, ns)
File "/global/project/projectdirs/acme/golaz/conda/envs/acme_diags_dev/lib/python2.7/site-packages/pkg_resources/__init__.py", line 1517, in run_script
exec(code, namespace, namespace)
File "/global/project/projectdirs/acme/golaz/conda/envs/acme_diags_dev/lib/python2.7/site-packages/acme_diags-1.0.0-py2.7.egg-info/scripts/acme_diags_driver.py", line 122, in <module>
parameters = cdp.cdp_run.serial(run_diag, parameters)
File "/global/project/projectdirs/acme/golaz/conda/envs/acme_diags_dev/lib/python2.7/site-packages/cdp/cdp_run.py", line 12, in serial
results.append(func(p))
File "/global/project/projectdirs/acme/golaz/conda/envs/acme_diags_dev/lib/python2.7/site-packages/acme_diags-1.0.0-py2.7.egg-info/scripts/acme_diags_driver.py", line 59, in run_diag
single_result = module.run_diag(parameters)
File "/global/project/projectdirs/acme/golaz/conda/envs/acme_diags_dev/lib/python2.7/site-packages/acme_diags/driver/lat_lon_driver.py", line 224, in run_diag
mv1_domain, diff, metrics_dict, parameter)
File "/global/project/projectdirs/acme/golaz/conda/envs/acme_diags_dev/lib/python2.7/site-packages/acme_diags/plot/__init__.py", line 43, in plot
plot_fcn(ref, test, diff, metrics_dict, parameter)
File "/global/project/projectdirs/acme/golaz/conda/envs/acme_diags_dev/lib/python2.7/site-packages/acme_diags/plot/cartopy/lat_lon_plot.py", line 149, in plot
(None, parameter.diff_title, None), parameter, stats=(max3, mean3, min3, r, c))
File "/global/project/projectdirs/acme/golaz/conda/envs/acme_diags_dev/lib/python2.7/site-packages/acme_diags/plot/cartopy/lat_lon_plot.py", line 61, in plot_panel
extend='both',
File "/global/project/projectdirs/acme/golaz/conda/envs/acme_diags_dev/lib/python2.7/site-packages/cartopy/mpl/geoaxes.py", line 1339, in contourf
if col.get_paths()])
File "/global/project/projectdirs/acme/golaz/conda/envs/acme_diags_dev/lib/python2.7/site-packages/matplotlib/collections.py", line 214, in get_datalim
paths = [transform.transform_path_non_affine(p) for p in paths]
File "/global/project/projectdirs/acme/golaz/conda/envs/acme_diags_dev/lib/python2.7/site-packages/matplotlib/transforms.py", line 2386, in transform_path_non_affine
return self._a.transform_path_non_affine(path)
File "/global/project/projectdirs/acme/golaz/conda/envs/acme_diags_dev/lib/python2.7/site-packages/cartopy/mpl/geoaxes.py", line 187, in transform_path_non_affine
geom, self.source_projection)
File "/global/project/projectdirs/acme/golaz/conda/envs/acme_diags_dev/lib/python2.7/site-packages/cartopy/crs.py", line 175, in project_geometry
return getattr(self, method_name)(geometry, src_crs)
File "/global/project/projectdirs/acme/golaz/conda/envs/acme_diags_dev/lib/python2.7/site-packages/cartopy/crs.py", line 330, in _project_polygon
return self._rings_to_multi_polygon(rings, is_ccw)
File "/global/project/projectdirs/acme/golaz/conda/envs/acme_diags_dev/lib/python2.7/site-packages/cartopy/crs.py", line 541, in _rings_to_multi_polygon
if prep_polygon.contains(interior_ring):
File "/global/project/projectdirs/acme/golaz/conda/envs/acme_diags_dev/lib/python2.7/site-packages/shapely/impl.py", line 37, in wrapper
return func(*args, **kwargs)
File "/global/project/projectdirs/acme/golaz/conda/envs/acme_diags_dev/lib/python2.7/site-packages/shapely/prepared.py", line 45, in contains
return bool(self.impl['prepared_contains'](self, other))
File "/global/project/projectdirs/acme/golaz/conda/envs/acme_diags_dev/lib/python2.7/site-packages/shapely/predicates.py", line 18, in __call__
self._check_topology(err, this, other)
File "/global/project/projectdirs/acme/golaz/conda/envs/acme_diags_dev/lib/python2.7/site-packages/shapely/topology.py", line 34, in _check_topology
if not geom.is_valid:
AttributeError: 'PreparedGeometry' object has no attribute 'is_valid'
@golaz You don't have the latest installed. In your stack trace, you have the following error:
File "/global/project/projectdirs/acme/golaz/conda/envs/acme_diags_dev/lib/python2.7/site-packages/acme_diags-1.0.0-py2.7.egg-info/scripts/acme_diags_driver.py", line 59, in run_diag
single_result = module.run_diag(parameters)
It says that on line 59 of acme_diags_driver.py, the line of code is single_result = module.run_diag(parameters). But look at line 59 of acme_diags_driver.py on master and you'll see it's a blank line. Hence, you don't have the latest code from master.
When you build the env from the acme_diags_env.yml (or even if you conda install it) it installs v1.0.0 of acme_diags from Anaconda. Then you install from source, but for whatever reason which Jill and I have experienced countless times, it does not install the latest from source.
So do:
rm -r /global/project/projectdirs/acme/golaz/conda/envs/acme_diags_dev/lib/python2.7/site-packages/acme_diags*
and reinstall from source. That should fix it.
I think the reason that you need to remove it is because when you install from source, there might be two different acme_diags in /path/to/your/env/lib/python2.7/site-packages/.
Ok, that could explain it. Please document the proper installation procedure on the documentation page:
https://acme-climate.github.io/acme_diags/docs/html/install-config-run.html
@golaz Yes, I'm getting to that now.
Updates: Strange thing!
I can't reproduce the error using a single script with the same data.
So we continued troubleshooting and think it might be problem from our end. We have a add_cyclic function to make up the last longitude, as follows:
def add_cyclic(var):
lon = var.getLongitude()
return var(longitude=(lon[0], lon[0] + 360.0, 'coe'))
When add_cyclic was commented out, no error. Plots all good (but you can notice the white gap at longitude 0W compared to the old plots).
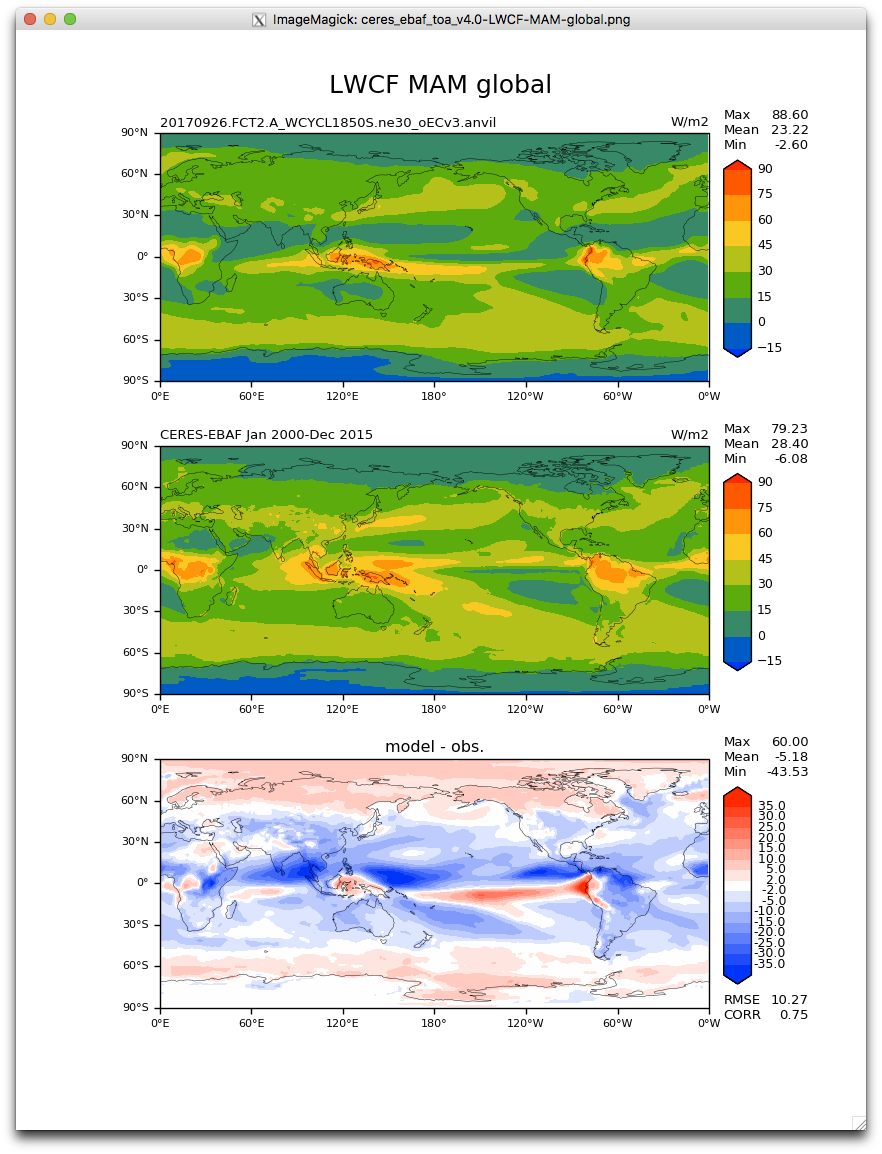 Now it starts to get mysterious...
Now it starts to get mysterious...
So it seems like this is not entirely a cartopy issue. But why does add_cyclic() only break for that certain variable, season, and contour levels? So for the most part, add_cyclic() with cartopy is okay.
This is quite strange. It might even be a bug related to cdms.
However, now that errors can be gracefully handled, this is not so urgent.
It is so random. It makes things more complicated with cdms envolved. The problem now becomes if we can reproduce this error without cdms to identify if this is a cartopy problem or cdms problem.
@chengzhuzhang, cartopy comes with its own version of add_cyclic, maybe you could try it out:
http://scitools.org.uk/cartopy/docs/v0.15/cartopy/util/util.html
There is also a matplotlib basemap one:
https://matplotlib.org/basemap/api/basemap_api.html
It probably would also be easy to do it with numpy.
Thank you, @golaz
I tried the add_cyclic_point from cartopy, the error remains. But I think I found the problem. It's mis-implementation from our side.
p1 = ax.contourf(lon, lat, var,
transform=proj,#ccrs.PlateCarree(),
norm=norm,
levels=levels,
cmap=cmap,
extend='both',
)
where,
proj = ccrs.PlateCarree(central_longitude=180)
See commit https://github.com/ACME-Climate/acme_diags/commit/0b1644ba691ecafd699031773aad61067a5686ed
I will try a full lat_lon run to see if this impact other diags
I ran the full lat_lon, non of the seasons broken :)