pylance-release
 pylance-release copied to clipboard
pylance-release copied to clipboard
Support for doccer-style docstring expansion
Hello! I've come across some situations where I don't get the expected docstrings in packages that, it seems, are using SciPy's doccer (or derived / similar tools) to reduce docstring redundancy. Essentially, doccer allows re-using of (parts of) existing docstrings, so one doesn't have to copy & paste recurring sections to different functions, and can keep a single "source of truth".
For example, the docstring of mne. compute_proj_raw looks something like this (omissions by me are indicated by an ellipsis):
@verbose
def compute_proj_raw(raw, start=0, stop=None, duration=1, n_grad=2, n_mag=2,
n_eeg=0, reject=None, flat=None, n_jobs=1, meg='separate',
verbose=None):
"""Compute SSP (spatial space projection) vectors on Raw.
Parameters
----------
raw : instance of Raw
A raw object to use the data from.
...
%(n_jobs)s
Number of jobs to use to compute covariance.
...
%(verbose)s
Returns
-------
projs: list
List of projection vectors.
See Also
--------
compute_proj_epochs, compute_proj_evoked
"""
n_jobs and verbose are defined in a central "documentation dictionary". When Sphinx builds the documentation, these placeholders are expanded.
To my surprise, the Python as well as the IPython REPL seem to be aware of this, and display the expanded docstrings when calling help(mne.compute_proj_raw):

In VS Code with Pylance, I receive the non-expanded docstrings, which are not really helpful:

Now I did a little bit of research, but couldn't really figure out just how Python and IPython generate the correct docstrings on the fly; but obviously it is possible somehow.
This is a feature I would very, very much like to see in Pylance as well. The current limitation extends beyond the MNE package I used in my example, and is likely to affect other scientific software packages as well, including SciPy.
Here, for completeness, is an example straight from a central SciPy function, namely ndimage.filters.correlate1d()
Function signature & docstring:
@_ni_docstrings.docfiller
def correlate1d(input, weights, axis=-1, output=None, mode="reflect",
cval=0.0, origin=0):
"""Calculate a 1-D correlation along the given axis.
The lines of the array along the given axis are correlated with the
given weights.
Parameters
----------
%(input)s
weights : array
1-D sequence of numbers.
%(axis)s
%(output)s
%(mode)s
%(cval)s
%(origin)s
Examples
--------
>>> from scipy.ndimage import correlate1d
>>> correlate1d([2, 8, 0, 4, 1, 9, 9, 0], weights=[1, 3])
array([ 8, 26, 8, 12, 7, 28, 36, 9])
"""
(truncated) output of help(scipy.ndimage.filters.correlate1d) on the Python 3.8 REPL

Pylance in VS Code
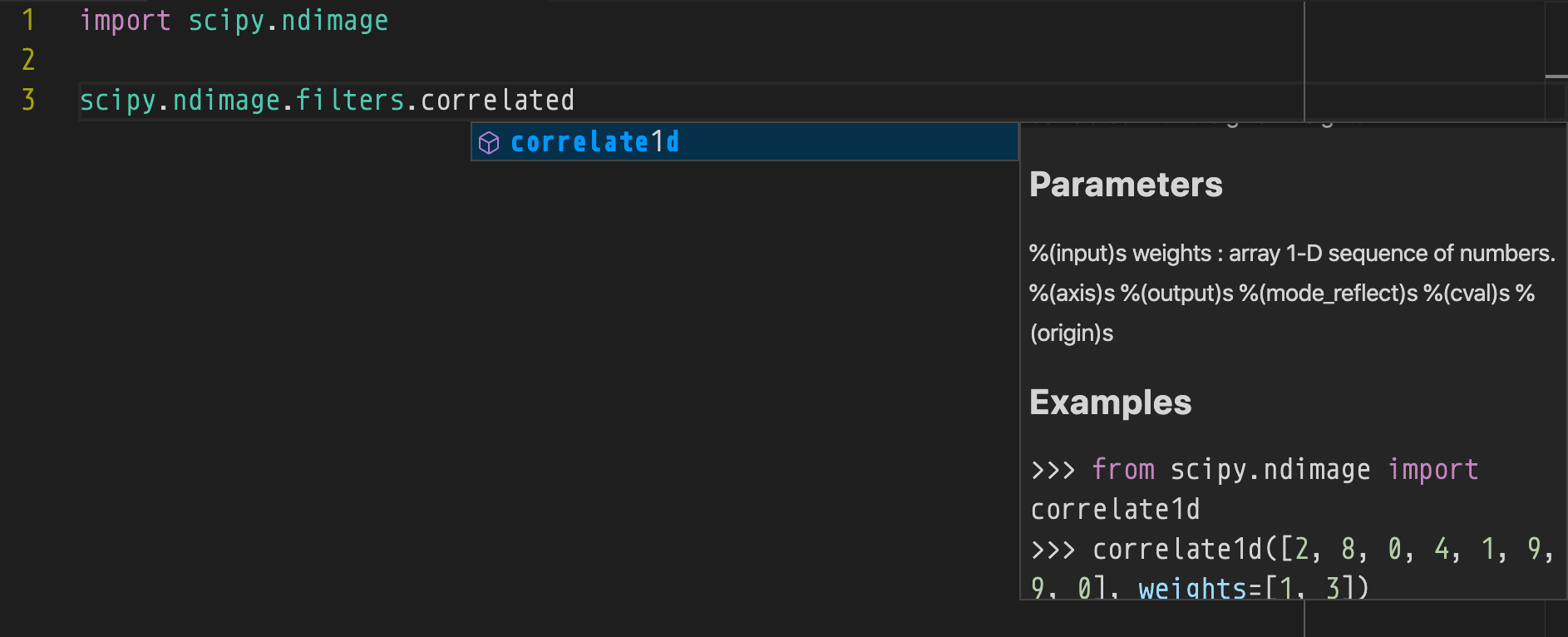
Even without expansion this displays incorrectly....
def add(self, item: SectionConfigItem, order: int = 10, deferred: bool = False, debug: bool = False) -> None:
"""
Add configuration to the output we're going to generate.
Parameters
----------
item : SectionConfigItem
Configuration to add, either a string, list of strings or another section.
order : int
Ordering of the item to add. Defaults to `10`.
deferred : bool
If the config item is a SectionBase, then rendering can be deferred until after configuration.
This is needed for instance for the constants section, which other protocols may add to.
debug : bool
The configuration provided should only be output when we are in debug mode.
"""
Pylance in VS Code...

@nkukard
Even without expansion this displays incorrectly....
Could be nicer, but I think it's a different issue – one of general docstring rendering :)
Newest improvements to docstrings about to be released.
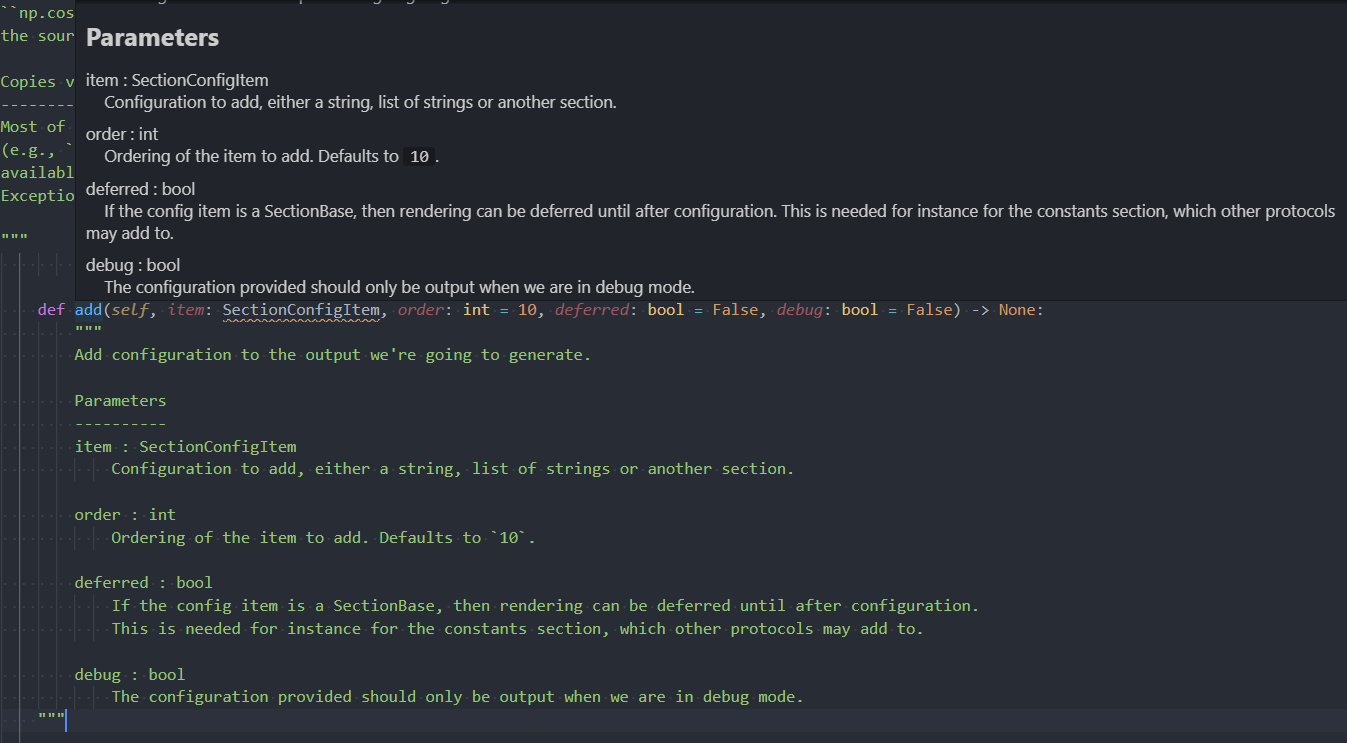
For reference, though, we didn't add this "expansion" that would normally happen at runtime. Not sure that we ever will given the non-static nature of these types of docstrings, but we can at least not print these indented regions poorly.
@jakebailey Thank you! Docstrings render much nicer now, great job!
I totally understand your argument regarding Pylance being a static type checker.
However, since these doccer expansions are used in a few important scientific packages (like SciPy), I'm afraid quite a bunch of users are affected by this. Maybe it's worth considering special-casing these types of docstrings? All that would need to be executed is the Python file defining the "doc dict", containing the strings to insert into the docstrings. This only needs to be read once, and the respective docstrings can be compiled either immediately or on-demand (at runtime), as string replacements should be super fast. 🙂
I'm not sure this is the same issue, but at least for MNE-Python I get very strange docstrings when hovering over a method (in the example, I'm using mne.time_frequency.tfr_multitaper source).
Here's the output when using Pylance:

And here's the (correct) output when using Jedi:
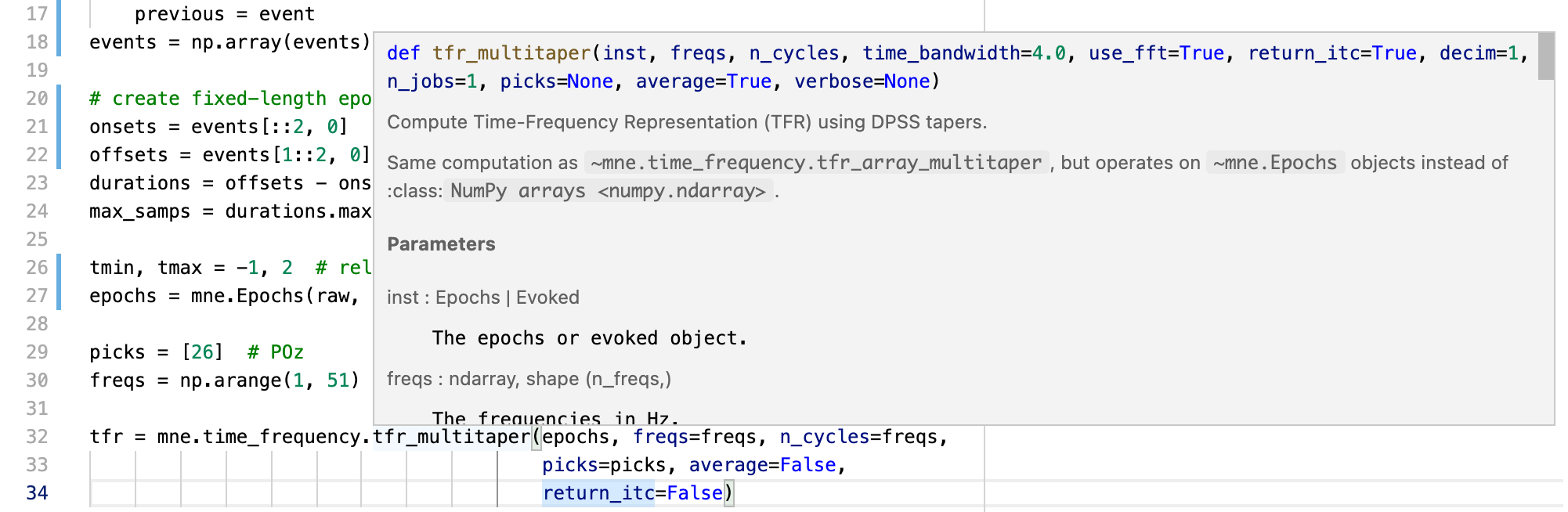
@cbrnr This is something else, related to the @verbose decorator used in MNE-Python. Care to open a separate issue for this one?
Are we sure it's not our fault? The verbose function doesn't use the functools.wraps decorator, although it seems like we're using decorator.FunctionMaker instead, which should do the same thing. I currently don't have time, I'll just switch back to Jedi and/or PyCharm. I just commented in case this was the same issue. But feel free to create a new issue if you can reproduce this and if it's not our fault.
see point 3 in https://github.com/microsoft/pyright/blob/master/docs/type-stubs.md#cleaning-up-generated-type-stubs for tips on how to add types to decorators.
Thank you, @bschnurr, I will try this
Hello again, and sorry for bumping this once more. But I'd like to once again bring to your attention that the lack of this feature is a real usability issue for numerous scientific / data science developers and users:
@hoechenberger
However, since these doccer expansions are used in a few important scientific packages (like SciPy), I'm afraid quite a bunch of users are affected by this. Maybe it's worth considering special-casing these types of docstrings? All that would need to be executed is the Python file defining the "doc dict", containing the strings to insert into the docstrings. This only needs to be read once, and the respective docstrings can be compiled either immediately or on-demand (at runtime), as string replacements should be super fast. 🙂
If this is not possible, do you have any recommendations as to what could be done on the side of the affected software projects to provide a better experience for Pylance users? While I know, for example, that type stubs can be a solution for libraries that do not yet use type annotations, could we provide something like "documentation stubs" that contain the expanded docstrings? The affected projects could do the expansion themselves and Pylance would just need to gather these strings. WDYT?
I'm not sure about "documentation stubs" (that requires greater thought, especially as the python ecosystem moves to inlining types as a single source of truth), but I do know that if regular type stubs are present and contain docs, we will use them and not try and analyze the real code.
but I do know that if regular type stubs are present and contain docs, we will use them and not try and analyze the real code.
Perfect, this could be something I could use, then. I will look into this.
Just note that type stubs effectively replace the source files in the eyes of the analysis, so if you add stubs, they should have all of the types filled out (as we're not going to go analyze the real code for anything).
Closing old issue. If this is still a problem, please reopen with the information requested. thanks
Hello @judej,
This is still topical and nothing has changed about this. Could you please reopen? This is still a big problem for those who use SciPy and certain other scientific tools. Thank you!
Thanks @judej!
one way to fix this would be to fixup stubs here https://github.com/microsoft/python-type-stubs/tree/main/scipy-stubs
Yes, I'd be thinking about this too 🤔
Closing for now... maybe there is something we can do with sphinx html generated docstrings