cbioportal-frontend
 cbioportal-frontend copied to clipboard
cbioportal-frontend copied to clipboard
Filter studies by reference genome
Allow filtering by reference genome when searching for studies on the landing page. The current search filter setup can easily be extended with additional study fields. Implementation of RFC 63. Work in progress.
- Add a search filter that allows filtering on one or more reference genomes (e.g.
reference-genome:hg19,hg38) - Add a graphical interface to 'toggle' the reference genomes that are available in the loaded studies:
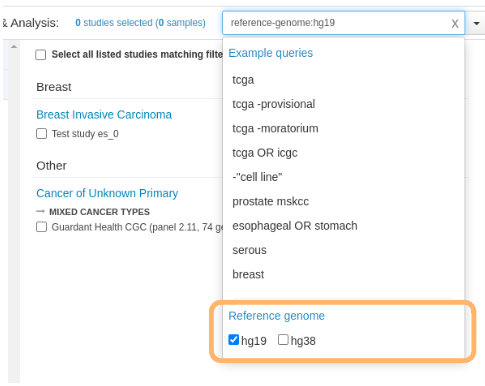
TODO:
- Add end to end tests
Changes
- Extended study search with a new filter syntax (i.e.
<prefix>:<value>) in which value can be a comma separated list - Create
SearchClauseandPhraseinterfaces to simplify search query modifications - Replace external autocomplete package with a minimal bootstrap
StudySearchsearch box component - Add dropdown form to StudySearch box that renders filters as defined in QueryParser
- Add
QueryParserthat contains a extendable list of search filter (see example below)- search prefix (e.g.
reference-genome) - relevant study fields (e.g.
study.referenceGenome) - rendering in drop down form (e.g. as a checkbox)
- search prefix (e.g.
Example search filter:
{
phrasePrefix: 'reference-genome',
nodeFields: ['referenceGenome'],
form: {
input: FilterCheckbox,
options: [...referenceGenomes],
label: 'Reference genome',
},
}
Checks
- Unit tests:
- Phrase.spec.tsx
- SearchClause.spec.tsx
- CheckboxFilterField.spec.tsx
- QueryParser.spec.ts
- textQueryUtils.spec.ts
- End to end tests:
- TODO
- Manual:
- open index page;
- start typing in search box;
- or open dropdown filter form
@BasLee did some manual testing of the feature. Seems good but for a couple things:
-
In the production version, the example query dropdown opens on focus and I think that's important behavior b/c it informs the user. I know this was a specific request of the original product designers. The behavior is a little nuanced. I think it only opens on focus IF the field is empty.
-
Escape key closes the menu. Probably not crucial, but nice.
-
Can you put a colon after the "Example queries" title? "Example queries:"
-
Can you nix the line beneath the example queries when there are no controls below it.
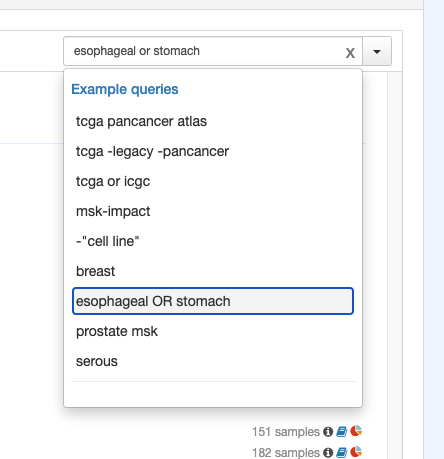
@BasLee did some manual testing of the feature. Seems good but for a couple things: [..]
@alisman, @pvannierop : all these things were solved in 889494c, 231e9ed and 471a8ab I guess?
Nice addition @BasLee !
I can't test the actual functionality in the public portal, but as @inodb suggested, the separator should not be present if there are no reference genome options.
In fact, I just agree with everything @inodb said :)
In terms of keeping 'reference-genome:' in the search bar - it's a really good point it takes up a lot of space that doesn't really exist. @inodb were you proposing to just show 'hg19' rather than 'reference-genome:hg19'? I worry that won't scale well - imagine a filter for 'has-mRNA:True' - just showing 'True' wouldn't be informative. I could go both ways at this point, but maybe it's ok for now, and we just need to think about a more general solution as we expand the feature beyond reference genome.
@inodb and @tmazor Thank you for reviewing! I think we already addressed the horizontal bar issue: will look into it. The search bar now shows a filter name and value, a 'gmail-like' syntax to allow filtering on specific fields, as described in the RFC and suggested by JJ and Sjoerd. The search bar might be a bit small for larger queries: as the number of filter options increases, we might want to give the search bar a more prominent place on the index page?
Thanks so much @BasLee! I like the wider search bar!
One more bug I found:
- [ ] Something broken with example filtering. Production:
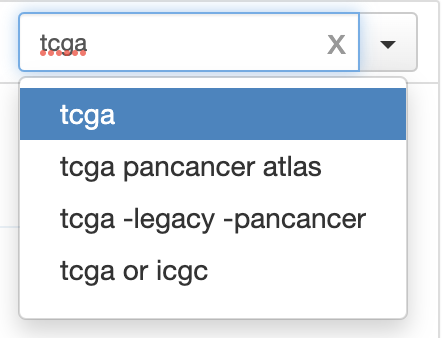 This PR keeps showing same examples:
This PR keeps showing same examples:
@tmazor: In terms of keeping 'reference-genome:' in the search bar - it's a really good point it takes up a lot of space that doesn't really exist. @inodb were you proposing to just show 'hg19' rather than 'reference-genome:hg19'? I worry that won't scale well - imagine a filter for 'has-mRNA:True' - just showing 'True' wouldn't be informative. I could go both ways at this point, but maybe it's ok for now, and we just need to think about a more general solution as we expand the feature beyond reference genome.
My thought was to remove it entirely (so no hg19 nor reference-genome: in the text field), only show the checkbox status in the dropdown, but I'm happy with the bigger search box as a solution for now. Thanks @BasLee ! I think the study text filter query language should eventually work in harmony with OQL
@BasLee one more question: is there a use case to be able to select both hg19 + hg38? If not it might be cleaner to just list in the examples:
reference-genome:hg19
reference-genome:hg38
then it would also filter nicely when typing reference-...
@BasLee one more question: is there a use case to be able to select both hg19 + hg38? If not it might be cleaner to just list in the examples:
reference-genome:hg19 reference-genome:hg38then it would also filter nicely when typing
reference-...
@inodb I think you want to show all studies by default? So that means selecting both hg19 and hg38 (when present)
One more bug I found:
@inodb: fixed in 9db3773
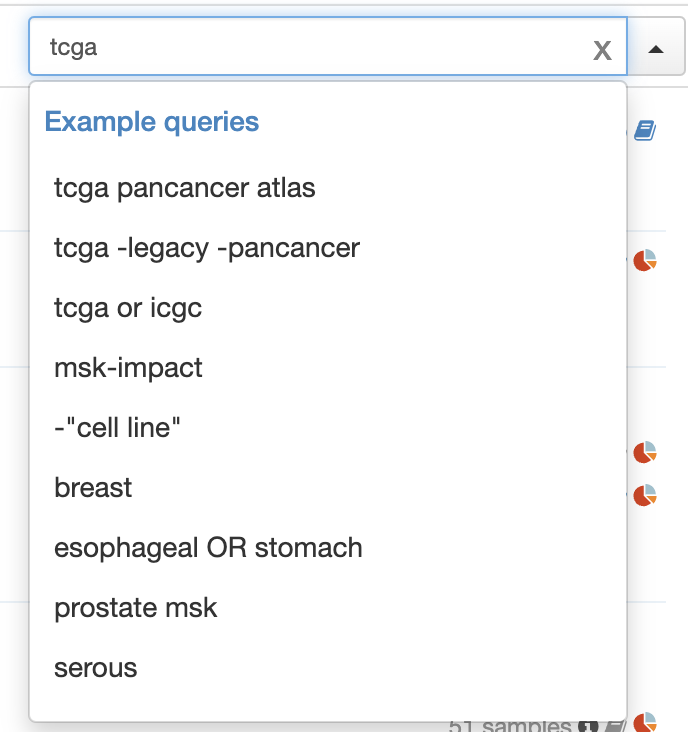
Thanks @BasLee ! Looks like there's still an issue vs production for queries not part of example queries:
PR:
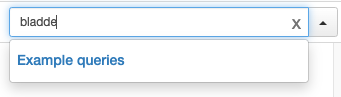
Production:
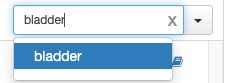
I think these should look identical
Note tho: the production behavior is also weird. Like it's kind odd that it only filters example queries. We can follow up with that later, but i think this PR should not introduce changes to current production behavior beyond adding the reference-genome filtering
Looks like there's still an issue vs production for queries not part of example queries [..]
@inodb a bit late, reading this just now... But yes, lets discuss what is should look like, with a follow up PR!