Error displaying MMTF structures in Jmol
When trying to display a structure in Jmol, for example from the MultipleAlignmentGUI, the following error gets thrown and the JmolPanel remains empty:
java.lang.ArithmeticException: / by zero at org.jmol.adapter.readers.cif.MessagePackReader.getInts(MessagePackReader.java:406) at org.jmol.adapter.readers.cif.MessagePackReader.decode(MessagePackReader.java:335) at org.jmol.adapter.readers.cif.MMTFReader.decode(MMTFReader.java:488) at org.jmol.adapter.readers.cif.MMTFReader.processBinaryDocument(MMTFReader.java:147) at org.jmol.adapter.smarter.AtomSetCollectionReader.readData(AtomSetCollectionReader.java:261) at org.jmol.adapter.smarter.SmarterJmolAdapter.staticGetAtomSetCollection(SmarterJmolAdapter.java:149) at org.jmol.adapter.smarter.SmarterJmolAdapter.getAtomSetCollection(SmarterJmolAdapter.java:142) at org.jmol.io.FileReader.run(FileReader.java:143) at org.jmol.viewer.FileManager.createAtomSetCollectionFromReader(FileManager.java:379) at org.jmol.viewer.Viewer.loadModelFromFile(Viewer.java:1697) at org.jmol.viewer.Viewer.loadModelFromFileRepaint(Viewer.java:1611) at org.jmol.viewer.Viewer.openReader(Viewer.java:1580) at org.jmol.api.JmolViewer.openReader(JmolViewer.java:319) at org.biojava.nbio.structure.align.gui.jmol.JmolPanel.setStructure(JmolPanel.java:146) at org.biojava.nbio.structure.align.gui.jmol.AbstractAlignmentJmol.setStructure(AbstractAlignmentJmol.java:159) at org.biojava.nbio.structure.align.gui.jmol.MultipleAlignmentJmol.initCoords(MultipleAlignmentJmol.java:358) at org.biojava.nbio.structure.align.gui.jmol.MultipleAlignmentJmol.(MultipleAlignmentJmol.java:342) at org.biojava.nbio.structure.align.gui.MultipleAlignmentJmolDisplay.display(MultipleAlignmentJmolDisplay.java:192) at org.biojava.nbio.structure.align.gui.MultipleAlignmentCalc.run(MultipleAlignmentCalc.java:90) at java.lang.Thread.run(Thread.java:745)
To reproduce the error, run the MultipleAlignmentGUI class with the provided example structures in the GUI.
Could that be a bug in jmol's decoder? The mmtf file is an original one produced by RCSB, isn't it?
Could that be a bug in jmol's decoder?
Yes, it is a problem in Jmol decoder, because opening an MMTF file in a simple JmolPanel shows the same error.
The mmtf file is an original one produced by RCSB, isn't it?
Yes, it should be fine... I am wondering if it can be that some of the changes between MMTF versions might have broken the MMTF decoder of Jmol. Reviewing the history of changes I see this:
- We updated MMTF to version 1.0.0: #563
- We updated the Jmol version and used mmCif to transmit structures: #569 (worked fine)
- We switched to MMTF for structure transmission to Jmol: #570 (do you remember if it worked ok? @sbliven )
- We updated MMTF to version 1.0.5 #646 (current - not working)
The possible solutions are:
- Revert #570 and use mmCif for transmission.
- Update the Jmol version if the bug has been patched.
Jmol's mmtf example page (V 14.20.4) seems to work: https://chemapps.stolaf.edu/jmol/jsmol/mmtf.htm
On Tue, Aug 8, 2017 at 3:30 AM, Aleix Lafita [email protected] wrote:
Could that be a bug in jmol's decoder?
Yes, it is a problem in Jmol decoder, because opening an MMTF file in a simple JmolPanel shows the same error.
The mmtf file is an original one produced by RCSB, isn't it?
Yes, it should be fine... I am wondering if it can be that some of the changes between MMTF versions might have broken the MMTF decoder of Jmol. Reviewing the history of changes I see this:
- We updated MMTF to version 1.0.0: #563 https://github.com/biojava/biojava/pull/563
- We updated the Jmol version and used mmCif to transmit structures: #569 https://github.com/biojava/biojava/pull/569 (worked fine)
- We switched to MMTF for structure transmission to Jmol: #570 https://github.com/biojava/biojava/pull/570 (do you remember if it worked ok? @sbliven https://github.com/sbliven )
- We updated MMTF to version 1.0.5 #646 https://github.com/biojava/biojava/pull/646 (current - not working)
The possible solutions are:
- Revert #570 https://github.com/biojava/biojava/pull/570 and use mmCif for transmission.
- Update the Jmol version if the bug has been patched.
— You are receiving this because you are subscribed to this thread. Reply to this email directly, view it on GitHub https://github.com/biojava/biojava/issues/629#issuecomment-320916875, or mute the thread https://github.com/notifications/unsubscribe-auth/ADuwEKmOIddXOVFmnTKWr8LNfQuutPaqks5sWDi5gaJpZM4Li-b7 .
We are using version 14.6.2 in BioJava, but strangely when I download the desktop version (same as the one in BioJava) showing an MMTF structure works, but when I try the same structure (same file) in the BioJava JmolPanel it shows the error.
I updated to Jmol 14.29.17 and the error is still there. I will submit a PR reverting to using mmCIF until we figure out what is the problem. For the purposes of visualizing structures the two formats are identical (except for little performance improvement).
The current error looks like this on the Jmol viewer, in case this helps:
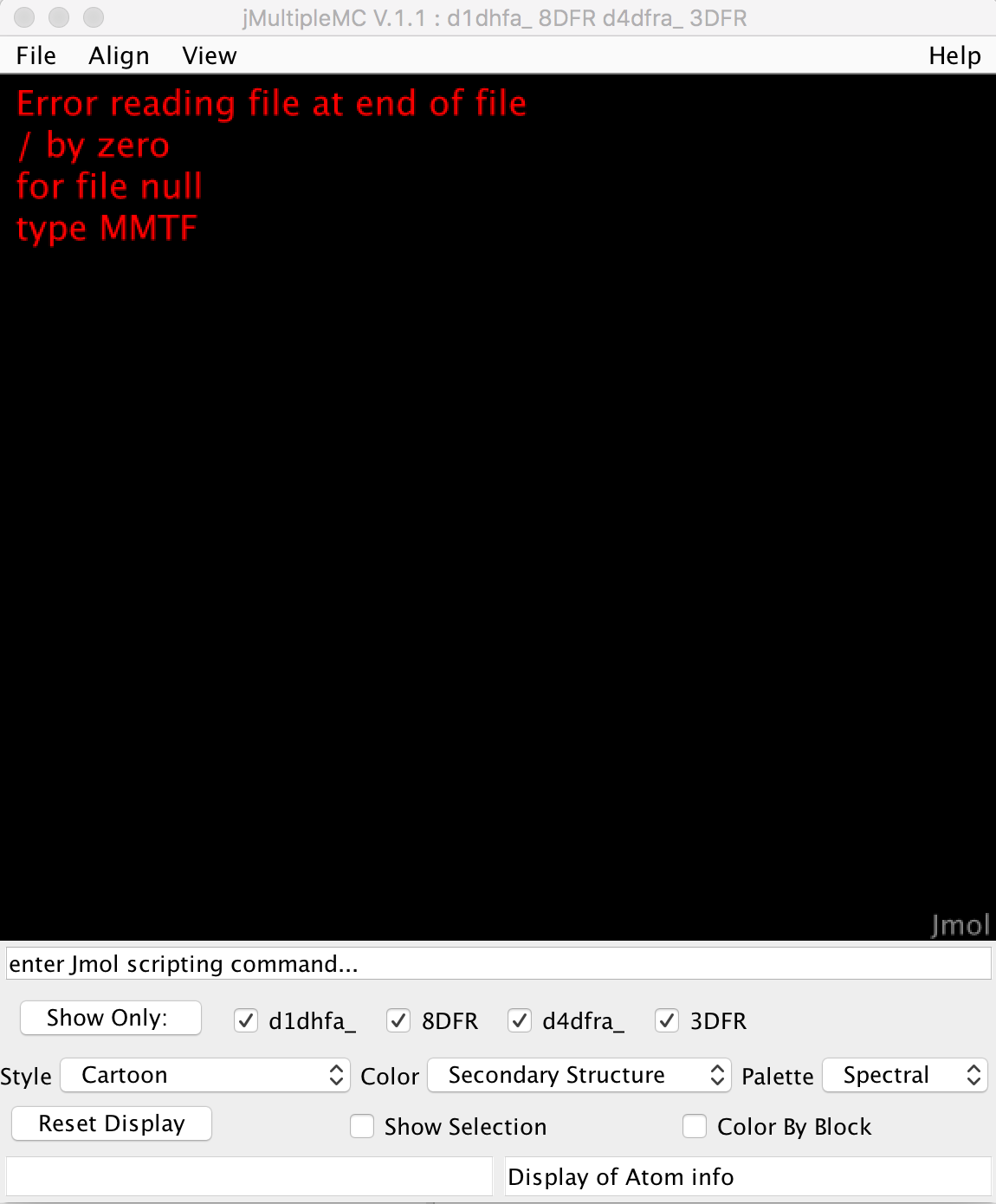
I did a bit of research and the problem has to be when converting the MMTF object into an OutputStream.
https://github.com/biojava/biojava/blob/359dce819e7678879ae41301b11fa268f92e81d1/biojava-structure-gui/src/main/java/org/biojava/nbio/structure/align/gui/jmol/JmolPanel.java#L130-L153
A possible test could be to save the OutputStream to a file and try opening it with a viewer.